

| Basic Information | |
|---|---|
| Species | Citrus clementina |
| Cazyme ID | Ciclev10017784m |
| Family | GT1 |
| Protein Properties | Length: 459 Molecular Weight: 50910.9 Isoelectric Point: 6.7153 |
| Chromosome | Chromosome/Scaffold: 2 Start: 33852535 End: 33853911 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 225 | 431 | 6.4e-24 |
| CDYIGSQFGKPVILSGPALPESPRLALEERWETLLGSFKSKSLIFCAFGSECVLNKEQFQELVLGFELSGLPFLVALKPPVGHDTIESALPEGFEERVKG RGFIHGGWVQQQLILKHPSVGCFVTHCGSGSLSEAMVNECQLVLLPNVGDQIINARLMGEELKVGVEVEKGDEDGLFTRDGVCKAVKAVIDDDHSEVGKE IKENHAK | |||
| Full Sequence |
|---|
| Protein Sequence Length: 459 Download |
| MGSETFHIAM YPWFAMGHLT PFLHIANKLA ERGHRISFLL PAKAITKFEP SNLHRDLITF 60 IPVSVPRVDG LPPGAETTND VPFPLHPLLM TAMDLTEPAI ESVLRRLKPD FVFFDFTHWL 120 PPLARKFGIK SVLYCIISPA TIGYLLSPER KLRERTLTDN DLLRPPQGFP TSKIRLRAHE 180 ARGLAAATVK EFGGGLSFAK RNLLSLSECD AIGFKTCREI EGPYCDYIGS QFGKPVILSG 240 PALPESPRLA LEERWETLLG SFKSKSLIFC AFGSECVLNK EQFQELVLGF ELSGLPFLVA 300 LKPPVGHDTI ESALPEGFEE RVKGRGFIHG GWVQQQLILK HPSVGCFVTH CGSGSLSEAM 360 VNECQLVLLP NVGDQIINAR LMGEELKVGV EVEKGDEDGL FTRDGVCKAV KAVIDDDHSE 420 VGKEIKENHA KWREFLRSER LENSHLDGFV QKLHGLLN* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02863 | PLN02863 | 5.0e-39 | 1 | 456 | 493 | + UDP-glucoronosyl/UDP-glucosyl transferase family protein | ||
| PLN02670 | PLN02670 | 8.0e-77 | 4 | 427 | 443 | + transferase, transferring glycosyl groups | ||
| PLN00414 | PLN00414 | 4.0e-143 | 1 | 458 | 463 | + glycosyltransferase family protein | ||
| PLN02208 | PLN02208 | 1.0e-151 | 5 | 454 | 455 | + glycosyltransferase family protein | ||
| PLN02764 | PLN02764 | 2.0e-152 | 1 | 458 | 460 | + glycosyltransferase family protein | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABL74480.1 | 0 | 1 | 457 | 1 | 458 | glucosyltransferase [Ipomoea batatas] |
| DDBJ | BAD95881.1 | 0 | 1 | 457 | 1 | 458 | glucosyltransferase [Ipomoea nil] |
| DDBJ | BAD95882.1 | 0 | 1 | 457 | 1 | 458 | glucosyltransferase [Ipomoea purpurea] |
| RefSeq | XP_002331944.1 | 0 | 10 | 457 | 1 | 445 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002522508.1 | 0 | 1 | 457 | 1 | 456 | UDP-glucosyltransferase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2vg8_A | 8e-22 | 266 | 434 | 269 | 444 | A Chain A, Ferric Horseradish Peroxidase C1a In Complex With Formate |
| PDB | 2vch_A | 8e-22 | 266 | 434 | 269 | 444 | A Chain A, Ferric Horseradish Peroxidase C1a In Complex With Formate |
| PDB | 2vce_A | 8e-22 | 266 | 434 | 269 | 444 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DY272554 | 336 | 1 | 336 | 0 |
| CB292845 | 299 | 19 | 317 | 0 |
| HO778286 | 429 | 32 | 459 | 0 |
| FC906269 | 287 | 1 | 287 | 0 |
| HO778286 | 24 | 9 | 32 | 0.0005 |
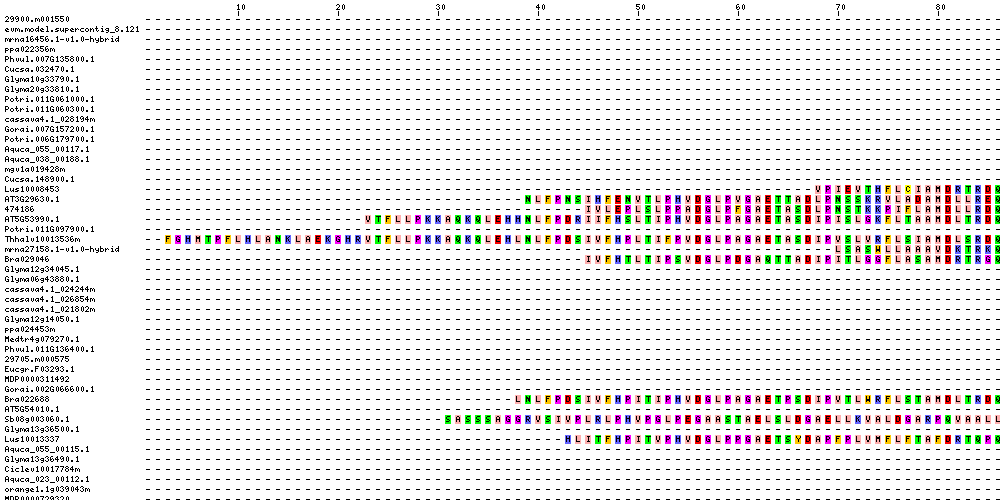
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |