

| Basic Information | |
|---|---|
| Species | Zea mays |
| Cazyme ID | GRMZM2G369803_T01 |
| Family | AA7 |
| Protein Properties | Length: 544 Molecular Weight: 58875.7 Isoelectric Point: 5.0895 |
| Chromosome | Chromosome/Scaffold: 6 Start: 105500352 End: 105502296 |
| Description | FAD-binding Berberine family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA7 | 71 | 314 | 0 |
| SATVRPLCIVTPVDASHVQAAVRCGRASGVRLRVRSGGHDYEGLSYRSERPEVFGVVDLSNLRAITVSADDDERPVPPTAPSAWVDSGATLGELYYTVAK NNPELAFPAGICPTIGVGGHLSGGGIGMMMRRFGLSVDNVLDAKLVNASGDLVDRAAMGEDHFWAIRGGGGESFGVVVSWKVGLVKVPSTVTAFNIVKTV ADQGAVDALTKWQDVAPGLPTDITIRVIIQGQRATFQSLYLGSC | |||
| Full Sequence |
|---|
| Protein Sequence Length: 544 Download |
| MAVLRGRLTL VLILSLNCCF SFPTVLSSVT SDGFLQCLSD NIPVGLIYTQ GSSNFTDVLV 60 SSVRNPRLFT SATVRPLCIV TPVDASHVQA AVRCGRASGV RLRVRSGGHD YEGLSYRSER 120 PEVFGVVDLS NLRAITVSAD DDERPVPPTA PSAWVDSGAT LGELYYTVAK NNPELAFPAG 180 ICPTIGVGGH LSGGGIGMMM RRFGLSVDNV LDAKLVNASG DLVDRAAMGE DHFWAIRGGG 240 GESFGVVVSW KVGLVKVPST VTAFNIVKTV ADQGAVDALT KWQDVAPGLP TDITIRVIIQ 300 GQRATFQSLY LGSCSDLVPV LNSSFPELGM TSADCLEMTW LESAAFFQFW NRRTPVEALL 360 DRKTSLSTFT KNKSDYVRRA IAKEAWESIF SWLTMDGAGM IILEPHGGFI GTVPDGATPY 420 PHRSGVLYNI QYITFWSAGG EQEGATATAW IGSFYEFMEQ HVSESPREAY VNYRDLDIGE 480 NVVVDDVSTL DSGRVWGEKY FAGNFQRLAA VKGVVDPTDY FRNEQSIPPQ STASRRRPGI 540 SLD* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
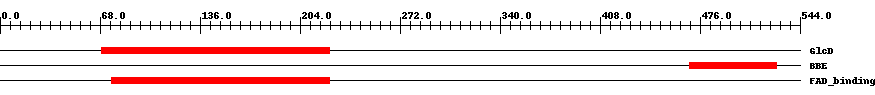 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0277 | GlcD | 2.0e-9 | 69 | 224 | 156 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| pfam08031 | BBE | 4.0e-16 | 469 | 528 | 60 | + Berberine and berberine like. This domain is found in the berberine bridge and berberine bridge- like enzymes which are involved in the biosynthesis of numerous isoquinoline alkaloids. They catalyze the transformation of the N-methyl group of (S)-reticuline into the C-8 berberine bridge carbon of (S)-scoulerine. | ||
| pfam01565 | FAD_binding_4 | 6.0e-17 | 76 | 224 | 149 | + FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidises the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAD53690.1 | 0 | 1 | 529 | 1 | 518 | putative CPRD2 [Oryza sativa Japonica Group] |
| GenBank | EAZ01289.1 | 0 | 26 | 529 | 26 | 521 | hypothetical protein OsI_23318 [Oryza sativa Indica Group] |
| GenBank | EEC80786.1 | 0 | 1 | 529 | 1 | 518 | hypothetical protein OsI_23315 [Oryza sativa Indica Group] |
| RefSeq | NP_001057830.1 | 0 | 40 | 529 | 82 | 561 | Os06g0549300 [Oryza sativa (japonica cultivar-group)] |
| RefSeq | XP_002438541.1 | 0 | 1 | 543 | 1 | 539 | hypothetical protein SORBIDRAFT_10g021685 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 4dns_B | 0 | 23 | 529 | 3 | 495 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 4dns_A | 0 | 23 | 529 | 3 | 495 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 3tsj_B | 0 | 31 | 529 | 8 | 495 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 3tsj_A | 0 | 31 | 529 | 8 | 495 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 3tsh_A | 0 | 31 | 529 | 8 | 495 | A Chain A, Crystal Structure Of Phl P 4, A Grass Pollen Allergen With Glucose Dehydrogenase Activity |