

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.006G234200.1 |
| Family | AA2 |
| Protein Properties | Length: 486 Molecular Weight: 53535.7 Isoelectric Point: 8.4946 |
| Chromosome | Chromosome/Scaffold: 06 Start: 48245753 End: 48252069 |
| Description | Peroxidase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA2 | 306 | 468 | 1.5e-35 |
| HREPRMGASLLRLHFHDCFVNVTSILHTTILCMPYNDLDELIFFVFQGCDGSLLLDSTSAFETEKNARGNFNSVRGFEVVDQIKAEVDRVCGRPVVSCAD ILAVAARDSVLALGGPTWKVRLGRRDSTTASRTLADSLLPSASMDLPALINNFKNQGLNKRDL | |||
| AA2 | 50 | 243 | 0 |
| REPRMGASLLRLHFHDCFVNVTSILHTTILCMPYNDLDELIFFVFQGCDGSLLLDSTSAFETEKNARGNFNSVRGFEVVDQIKAEVDRVCGRPVVSCADI LAVAARDSVLALGGPTWKVRLGRRDSTTASRTLADSLLPSASMDLPALINNFKNQGLNKRDLVALSGGHTIGLSQCVIFRNRIYNATNIDPAFA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 486 Download |
| MAPRFNFLLH AFLCLALATT SFSLSPKFYD NLCPQALPAI KRIVEAAVHR EPRMGASLLR 60 LHFHDCFVNV TSILHTTILC MPYNDLDELI FFVFQGCDGS LLLDSTSAFE TEKNARGNFN 120 SVRGFEVVDQ IKAEVDRVCG RPVVSCADIL AVAARDSVLA LGGPTWKVRL GRRDSTTASR 180 TLADSLLPSA SMDLPALINN FKNQGLNKRD LVALSGGHTI GLSQCVIFRN RIYNATNIDP 240 AFAKERRATC PRTGGNTNLA PFDPTPARFD TAYFKNLVKE RGLLTSDQAL FSALPAIKRI 300 VEAAVHREPR MGASLLRLHF HDCFVNVTSI LHTTILCMPY NDLDELIFFV FQGCDGSLLL 360 DSTSAFETEK NARGNFNSVR GFEVVDQIKA EVDRVCGRPV VSCADILAVA ARDSVLALGG 420 PTWKVRLGRR DSTTASRTLA DSLLPSASMD LPALINNFKN QGLNKRDLCS SLRWPHHRIV 480 TMRYL* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
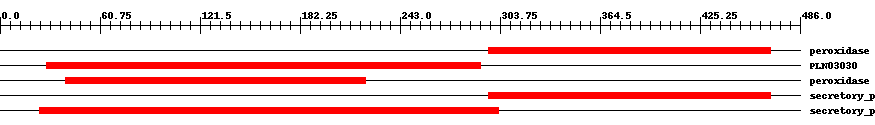 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00141 | peroxidase | 3.0e-48 | 297 | 468 | 172 | + Peroxidase. | ||
| PLN03030 | PLN03030 | 1.0e-52 | 28 | 292 | 274 | + cationic peroxidase; Provisional | ||
| pfam00141 | peroxidase | 6.0e-56 | 40 | 222 | 183 | + Peroxidase. | ||
| cd00693 | secretory_peroxidase | 1.0e-62 | 297 | 468 | 172 | + Horseradish peroxidase and related secretory plant peroxidases. Secretory peroxidases belong to class III of the plant heme-dependent peroxidase superfamily. All members of the superfamily share a heme prosthetic group and catalyze a multistep oxidative reaction involving hydrogen peroxide as the electron acceptor. Class III peroxidases are found in the extracellular space or in the vacuole in plants where they have been implicated in hydrogen peroxide detoxification, auxin catabolism and lignin biosynthesis, and stress response. Class III peroxidases contain four conserved disulphide bridges and two conserved calcium binding sites. | ||
| cd00693 | secretory_peroxidase | 9.0e-117 | 24 | 303 | 288 | + Horseradish peroxidase and related secretory plant peroxidases. Secretory peroxidases belong to class III of the plant heme-dependent peroxidase superfamily. All members of the superfamily share a heme prosthetic group and catalyze a multistep oxidative reaction involving hydrogen peroxide as the electron acceptor. Class III peroxidases are found in the extracellular space or in the vacuole in plants where they have been implicated in hydrogen peroxide detoxification, auxin catabolism and lignin biosynthesis, and stress response. Class III peroxidases contain four conserved disulphide bridges and two conserved calcium binding sites. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004601 | peroxidase activity |
| GO:0006979 | response to oxidative stress |
| GO:0020037 | heme binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002268412.1 | 0 | 1 | 293 | 1 | 269 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002268412.1 | 0 | 293 | 468 | 38 | 187 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002323055.1 | 0 | 24 | 293 | 1 | 244 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002531319.1 | 0 | 1 | 291 | 1 | 269 | Peroxidase 2 precursor, putative [Ricinus communis] |
| RefSeq | XP_002531319.1 | 0 | 287 | 468 | 33 | 188 | Peroxidase 2 precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1sch_B | 0 | 24 | 301 | 2 | 253 | A Chain A, Peanut Peroxidase |
| PDB | 1sch_B | 0 | 292 | 468 | 13 | 162 | A Chain A, Peanut Peroxidase |
| PDB | 1sch_A | 0 | 24 | 301 | 2 | 253 | A Chain A, Peanut Peroxidase |
| PDB | 1sch_A | 0 | 292 | 468 | 13 | 162 | A Chain A, Peanut Peroxidase |
| PDB | 7atj_A | 0 | 24 | 300 | 2 | 260 | A Chain A, Peanut Peroxidase |