

| Basic Information | |
|---|---|
| Species | Setaria italica |
| Cazyme ID | Si039501m |
| Family | GT1 |
| Protein Properties | Length: 481 Molecular Weight: 50313.3 Isoelectric Point: 6.1264 |
| Chromosome | Chromosome/Scaffold: 9 Start: 53138156 End: 53139598 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 282 | 464 | 3.2e-38 |
| RVLEWLDTKPVRSVVYVCFGSLTRFPREQVSELGMGLAYSGANFVWVVGDKSAPPLPDIDAAAPGRGLVVRGWAPQVAVLRHAAVGAFVTHCGWGAVTEA AAAGVPVLTWPVFAEQFYNEALVVGIAGTGVAMGAERGYVWGGEALGGVVVGREAVAERVRSALADKALRRRAGEIGGRARRA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 481 Download |
| MRPSSTSVAS SGDTAPPRMY FIPFPTPGHA LPMSDLARLF ASRGADATLV LTRANAARLG 60 GPVARAAAAG LRIRVHALTL PAEAAGLTGG HESADDLPNR ELAGPFAVAV DLLAPLFADL 120 LRRHPADAVV FDGVLPWAAT AAPELGIPRY AFTGTGCFAL SVQRALLLHS PQNGVASDTE 180 PFLVPGLPDP VRLTRSRLAE ATLPGADSRE FLNRMFDVER ATAGWVVNSF ADLEQRYVEH 240 YEKVTGKPVF AVGPVCFVNG DGDDTLERGR GGEAVAAAEA ARVLEWLDTK PVRSVVYVCF 300 GSLTRFPREQ VSELGMGLAY SGANFVWVVG DKSAPPLPDI DAAAPGRGLV VRGWAPQVAV 360 LRHAAVGAFV THCGWGAVTE AAAAGVPVLT WPVFAEQFYN EALVVGIAGT GVAMGAERGY 420 VWGGEALGGV VVGREAVAER VRSALADKAL RRRAGEIGGR ARRAVEAGGS SYEAVGALLE 480 D |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN03004 | PLN03004 | 8.0e-42 | 144 | 421 | 293 | + UDP-glycosyltransferase | ||
| PLN00164 | PLN00164 | 6.0e-44 | 15 | 418 | 441 | + glucosyltransferase; Provisional | ||
| PLN02863 | PLN02863 | 8.0e-63 | 23 | 405 | 399 | + UDP-glucoronosyl/UDP-glucosyl transferase family protein | ||
| PLN03007 | PLN03007 | 1.0e-91 | 21 | 418 | 412 | + UDP-glucosyltransferase family protein | ||
| PLN02534 | PLN02534 | 6.0e-101 | 15 | 422 | 422 | + UDP-glycosyltransferase | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABF94606.1 | 0 | 10 | 481 | 2 | 478 | Flavonol-3-O-glycoside-7-O-glucosyltransferase 1, putative, expressed [Oryza sativa (japonica cultivar-group)] |
| DDBJ | BAH92039.1 | 0 | 10 | 481 | 2 | 478 | Os03g0212000 [Oryza sativa Japonica Group] |
| GenBank | EAY89016.1 | 0 | 10 | 441 | 2 | 438 | hypothetical protein OsI_10499 [Oryza sativa Indica Group] |
| GenBank | EAZ26035.1 | 0 | 10 | 481 | 2 | 478 | hypothetical protein OsJ_09889 [Oryza sativa Japonica Group] |
| RefSeq | XP_002468294.1 | 0 | 1 | 481 | 1 | 481 | hypothetical protein SORBIDRAFT_01g043140 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2acv_B | 0 | 18 | 481 | 11 | 460 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 2acv_A | 0 | 18 | 481 | 11 | 460 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 2acw_B | 0 | 18 | 481 | 11 | 460 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2acw_A | 0 | 18 | 481 | 11 | 460 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2pq6_A | 8e-40 | 17 | 481 | 9 | 476 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
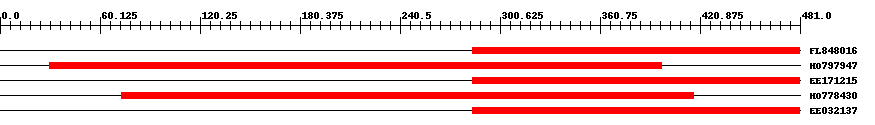 | ||||
| Hit | Length | Start | End | EValue |
| FL848016 | 198 | 284 | 481 | 0 |
| HO797947 | 376 | 30 | 398 | 0 |
| EE171215 | 198 | 284 | 481 | 0 |
| HO778430 | 352 | 73 | 417 | 0 |
| EE032137 | 198 | 284 | 481 | 0 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |