

| Basic Information | |
|---|---|
| Species | Mimulus guttatus |
| Cazyme ID | mgv1a027071m |
| Family | CBM22 |
| Protein Properties | Length: 897 Molecular Weight: 99807.7 Isoelectric Point: 5.9268 |
| Chromosome | Chromosome/Scaffold: 91 Start: 606529 End: 610744 |
| Description | glycosyl hydrolase family 10 protein / carbohydrate-binding domain-containing protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM22 | 14 | 153 | 5e-26 |
| NIIINHDFSGGLHLWHPNSCEAFLVSQETTRPKGLSDNLSAPFAVITKRKQQWQGLEQDITDRVSPFSVYNICAFVGISSGASGNQEHVLATLKLEFEDN SVRYLFIGRTCASTEHWEKVEGTFSLSAMPRRVVFYVEGP | |||
| CBM22 | 177 | 313 | 2.9e-29 |
| NIIQNPRFDDGLNNWSGRGCKIALHDTMSDGNILPVSGKFFGSTENRTDYWNGIQQDITGQVKRKLAYDFIATVRIFGNNISAANVKATLYIQTADLREQ YIGVASVQATDKDWVQLKGKFLVNGSPSRAVIFLEGP | |||
| CBM22 | 348 | 489 | 3.80032e-42 |
| NVIANSNLNDGTNGWFPLGNCNLSVGNGSPHILPPMAKDSLGAHEPLSGSYILVTNRTQTWMGPAQMITEKLKLYLTYQVSAWIRVANHASKPQNINIAL GVDGQWVNGGQIESSDDKWHEVGGSFRIEKQPTKVMVYVQGP | |||
| GH10 | 549 | 820 | 0 |
| QNTFPFGTCINRSNIDNEDIVDFFTKNFNWSVFENELKWYWTEPQKGNLNYKDADDLLNLCANHNIQLRGHCIFWEAESSVQSWIRNLNKDDLTSAVENR LAGLLARYNGKFKHYDVNNEMLHGSFYQDRLGKDIRAHMFKTAYQLDPTATLFVNDYNIEDGCDARSSPEKYIEHILDLRAQGGPVGGIGVQGHINSPVG PVVRSALDKLGVLGLPIWFTELDVASDNEFVRADDLEVMLRESFAHPAVEGVVLWGFWELFMSRDNAYLVNA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 897 Download |
| SPNNEPMDSH YSNNIIINHD FSGGLHLWHP NSCEAFLVSQ ETTRPKGLSD NLSAPFAVIT 60 KRKQQWQGLE QDITDRVSPF SVYNICAFVG ISSGASGNQE HVLATLKLEF EDNSVRYLFI 120 GRTCASTEHW EKVEGTFSLS AMPRRVVFYV EGPSPGVDLL VKSVVISCIS FSQCEENIIQ 180 NPRFDDGLNN WSGRGCKIAL HDTMSDGNIL PVSGKFFGST ENRTDYWNGI QQDITGQVKR 240 KLAYDFIATV RIFGNNISAA NVKATLYIQT ADLREQYIGV ASVQATDKDW VQLKGKFLVN 300 GSPSRAVIFL EGPPPGTDIL LDNLVVKHAA KAPPASPPVV ENAAFGVNVI ANSNLNDGTN 360 GWFPLGNCNL SVGNGSPHIL PPMAKDSLGA HEPLSGSYIL VTNRTQTWMG PAQMITEKLK 420 LYLTYQVSAW IRVANHASKP QNINIALGVD GQWVNGGQIE SSDDKWHEVG GSFRIEKQPT 480 KVMVYVQGPE AGVDLMVAGL QIFPVDRRAR FRQLKKETDL IRKRDVILKF SSSDSATLVG 540 TSVKIRQIQN TFPFGTCINR SNIDNEDIVD FFTKNFNWSV FENELKWYWT EPQKGNLNYK 600 DADDLLNLCA NHNIQLRGHC IFWEAESSVQ SWIRNLNKDD LTSAVENRLA GLLARYNGKF 660 KHYDVNNEML HGSFYQDRLG KDIRAHMFKT AYQLDPTATL FVNDYNIEDG CDARSSPEKY 720 IEHILDLRAQ GGPVGGIGVQ GHINSPVGPV VRSALDKLGV LGLPIWFTEL DVASDNEFVR 780 ADDLEVMLRE SFAHPAVEGV VLWGFWELFM SRDNAYLVNA EGDLNEAGKR YVALKQEWLS 840 RARGRIDEQG KFGFRGFYGV YEVEVEVAND VSKEKMVKAS FVVDKGEGPI VISINL* 900 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
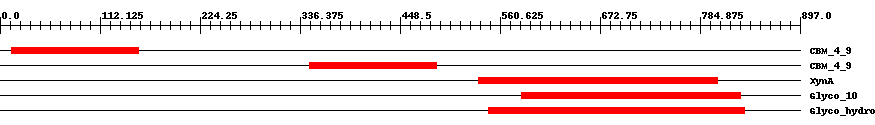 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam02018 | CBM_4_9 | 5.0e-15 | 13 | 155 | 144 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| pfam02018 | CBM_4_9 | 9.0e-16 | 347 | 489 | 149 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| COG3693 | XynA | 1.0e-29 | 536 | 805 | 291 | + Beta-1,4-xylanase [Carbohydrate transport and metabolism] | ||
| smart00633 | Glyco_10 | 1.0e-67 | 585 | 830 | 267 | + Glycosyl hydrolase family 10. | ||
| pfam00331 | Glyco_hydro_10 | 4.0e-71 | 548 | 835 | 308 | + Glycosyl hydrolase family 10. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| GO:0016798 | hydrolase activity, acting on glycosyl bonds |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAZ79233.1 | 0 | 3 | 896 | 21 | 918 | putative xylanase Xyn2 [Nicotiana tabacum] |
| RefSeq | XP_002301133.1 | 2e-30 | 3 | 326 | 324 | 660 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 7 | 896 | 155 | 1049 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 13 | 497 | 1 | 479 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 176 | 502 | 1 | 314 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1i1x_A | 5e-31 | 554 | 804 | 18 | 268 | A Chain A, Crystal Structural Analysis Of Active Site Mutant Q189e Of Lgtc |
| PDB | 1i1w_A | 5e-31 | 554 | 804 | 18 | 268 | A Chain A, 0.89a Ultra High Resolution Structure Of A Thermostable Xylanase From Thermoascus Aurantiacus |
| PDB | 2bnj_A | 9e-31 | 554 | 804 | 18 | 268 | A Chain A, The Xylanase Ta From Thermoascus Aurantiacus Utilizes Arabinose Decorations Of Xylan As Significant Substrate Specificity Determinants |
| PDB | 1tux_A | 3e-30 | 554 | 804 | 18 | 268 | A Chain A, High Resolution Crystal Structure Of A Thermostable Xylanase From Thermoascus Aurantiacus |
| PDB | 1k6a_A | 8e-30 | 554 | 804 | 18 | 268 | A Chain A, Structural Studies On The Mobility In The Active Site Of The Thermoascus Aurantiacus Xylanase I |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EB442796 | 298 | 375 | 672 | 0 |
| GO850973 | 328 | 473 | 800 | 0 |
| GR184067 | 268 | 554 | 821 | 0 |
| EB442796 | 120 | 47 | 166 | 0.00000004 |
| EB442796 | 114 | 211 | 324 | 0.00007 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |