

| Basic Information | |
|---|---|
| Species | Citrus sinensis |
| Cazyme ID | orange1.1g012467m |
| Family | AA1 |
| Protein Properties | Length: 464 Molecular Weight: 51656.6 Isoelectric Point: 7.7607 |
| Chromosome | Chromosome/Scaffold: 00028 Start: 935766 End: 939923 |
| Description | Plant L-ascorbate oxidase |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA1 | 8 | 435 | 0 |
| QAGTYFYHGHLGMQRSAGLYGSLIVDVADGEKEPFHYDGEFNLLLSDWWHRSVHEQEVGLSSRPLRWIGEPQTLLINGRGQFNCSLAAHFSNGSAEQCKL RGNEQCAPQILHVQPNKTYRLRIASTTALASLNLAVKNHKMVVVEADGNYVQPFEVDDMDIYSGESYSVLLTTNQDPSYNYWISAGVRGRKPATPPALTL LNYHPTSASKIPLSPPPITPRWDDYDHSKSFSNKIFALMGSPKPPTNFHRRLTLLNTQNTINGFTKWAINNVSLTLPPTPYLGSIKYGLKDAFDQNGPPE NFSNEYDVMKPPVNANTTLGSGVYMLGLNTTVDVILQNANAIRPNLSEIHPWHLHGHDFWVLGRGEGKFTKEDEKKFNLKNPPLKNTAVIFPYGWTALRF VADNPGAWAFHCHIEPHFHIGMGVVLAL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 464 Download |
| MLDVWQVQAG TYFYHGHLGM QRSAGLYGSL IVDVADGEKE PFHYDGEFNL LLSDWWHRSV 60 HEQEVGLSSR PLRWIGEPQT LLINGRGQFN CSLAAHFSNG SAEQCKLRGN EQCAPQILHV 120 QPNKTYRLRI ASTTALASLN LAVKNHKMVV VEADGNYVQP FEVDDMDIYS GESYSVLLTT 180 NQDPSYNYWI SAGVRGRKPA TPPALTLLNY HPTSASKIPL SPPPITPRWD DYDHSKSFSN 240 KIFALMGSPK PPTNFHRRLT LLNTQNTING FTKWAINNVS LTLPPTPYLG SIKYGLKDAF 300 DQNGPPENFS NEYDVMKPPV NANTTLGSGV YMLGLNTTVD VILQNANAIR PNLSEIHPWH 360 LHGHDFWVLG RGEGKFTKED EKKFNLKNPP LKNTAVIFPY GWTALRFVAD NPGAWAFHCH 420 IEPHFHIGMG VVLALGVETV GNIPNQALAC GLTGKRFMNP KQN* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR03390 | ascorbOXfungal | 6.0e-55 | 9 | 436 | 464 | + L-ascorbate oxidase, fungal type. This model describes a family of fungal ascorbate oxidases, within a larger family of multicopper oxidases that also includes plant ascorbate oxidases (TIGR03388), plant laccases and laccase-like proteins (TIGR03389), and related proteins. The member from Acremonium sp. HI-25 is characterized. | ||
| TIGR03389 | laccase | 1.0e-56 | 6 | 448 | 471 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| PLN02604 | PLN02604 | 8.0e-175 | 8 | 453 | 449 | + oxidoreductase | ||
| TIGR03388 | ascorbase | 0 | 8 | 453 | 447 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| PLN02191 | PLN02191 | 0 | 8 | 463 | 458 | + L-ascorbate oxidase | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005507 | copper ion binding |
| GO:0016491 | oxidoreductase activity |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABK96191.1 | 0 | 8 | 458 | 136 | 588 | unknown [Populus trichocarpa] |
| EMBL | CAN82127.1 | 0 | 8 | 459 | 102 | 553 | hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002275678.1 | 0 | 8 | 459 | 125 | 576 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002299782.1 | 0 | 8 | 458 | 108 | 560 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002312838.1 | 0 | 8 | 458 | 138 | 591 | l-ascorbate oxidase precursor [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1asq_B | 0 | 10 | 463 | 99 | 551 | A Chain A, The Structure Of A Protein In Glycosyl Transferase Family 8 From Anaerococcus Prevotii. |
| PDB | 1asq_A | 0 | 10 | 463 | 99 | 551 | A Chain A, The Structure Of A Protein In Glycosyl Transferase Family 8 From Anaerococcus Prevotii. |
| PDB | 1asp_B | 0 | 10 | 463 | 99 | 551 | A Chain A, The Structure Of A Protein In Glycosyl Transferase Family 8 From Anaerococcus Prevotii. |
| PDB | 1asp_A | 0 | 10 | 463 | 99 | 551 | A Chain A, The Structure Of A Protein In Glycosyl Transferase Family 8 From Anaerococcus Prevotii. |
| PDB | 1aso_B | 0 | 10 | 463 | 99 | 551 | A Chain A, The Structure Of A Protein In Glycosyl Transferase Family 8 From Anaerococcus Prevotii. |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DY305215 | 310 | 151 | 460 | 0 |
| FC929669 | 290 | 151 | 440 | 0 |
| CX044124 | 281 | 182 | 462 | 0 |
| CX046237 | 270 | 15 | 284 | 0 |
| DN799381 | 270 | 86 | 355 | 0 |
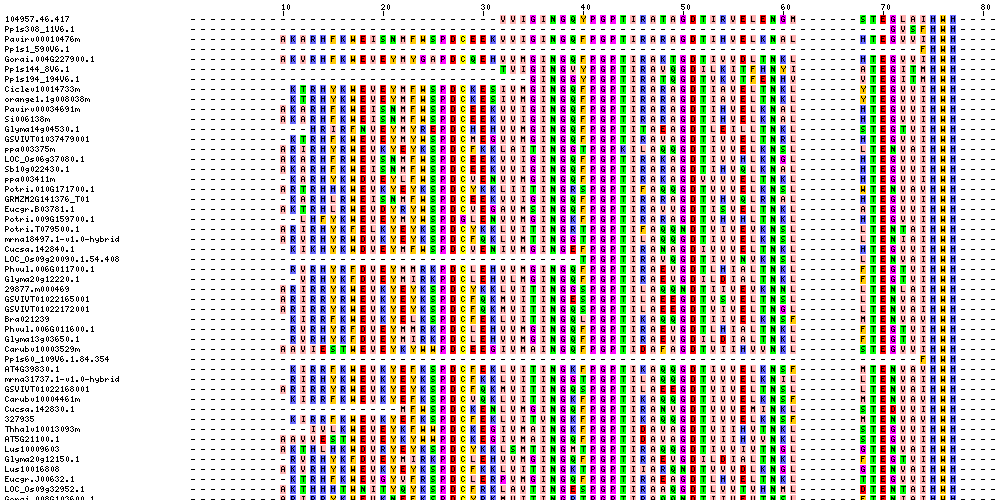
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |