You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004377_00291
You are here: Home > Sequence: MGYG000004377_00291
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-485; | |||||||||||
| CAZyme ID | MGYG000004377_00291 | |||||||||||
| CAZy Family | CBM67 | |||||||||||
| CAZyme Description | Beta-galactosidase BoGH2A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 35; End: 1108 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam18565 | Glyco_hydro2_C5 | 1.11e-41 | 251 | 353 | 1 | 103 | Glycoside hydrolase family 2 C-terminal domain 5. Domain 5 is found in dimeric beta-D-galactosidase from Paracoccus sp. 32d, which contributes to stabilization of the functional dimer. It is suggested that the location of this domain 5, may be one of the factors responsible for the creation of a functional dimer and cold-adaptation of this enzyme. |
| pfam16355 | DUF4982 | 6.91e-21 | 174 | 239 | 1 | 61 | Domain of unknown function (DUF4982). This family is found in the C-terminal of uncharacterized proteins and beta-galactosidases around 680 residues in length from various Bacteroides species. The function of this protein is unknown. |
| smart00634 | BID_1 | 1.66e-05 | 251 | 353 | 3 | 91 | Bacterial Ig-like domain (group 1). |
| pfam02369 | Big_1 | 2.08e-04 | 262 | 342 | 1 | 64 | Bacterial Ig-like domain (group 1). This family consists of bacterial domains with an Ig-like fold. Members of this family are found in bacterial surface proteins such as intimins and invasins involved in pathogenicity. |
| pfam09134 | Invasin_D3 | 4.84e-04 | 248 | 354 | 2 | 95 | Invasin, domain 3. Members of this family adopt a structure consisting of an immunoglobulin-like beta-sandwich, with seven strands in two beta-sheets, arranged in a Greek-key topology. It forms part of the extracellular region of the protein, which can be expressed as a soluble protein (Inv497) that binds integrins and promotes subsequent uptake by cells when attached to bacteria. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCD35275.1 | 6.31e-182 | 1 | 357 | 458 | 811 |
| QDM10520.1 | 5.64e-171 | 1 | 357 | 456 | 819 |
| QUT79099.1 | 5.64e-171 | 1 | 357 | 456 | 819 |
| QUR43922.1 | 6.58e-166 | 1 | 357 | 464 | 815 |
| QDH55004.1 | 9.31e-166 | 1 | 357 | 464 | 815 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5T98_A | 2.47e-90 | 9 | 357 | 459 | 821 | Crystalstructure of BuGH2Awt [Bacteroides uniformis],5T98_B Crystal structure of BuGH2Awt [Bacteroides uniformis],5T99_A Crystal structure of BuGH2Awt in complex with Galactoisofagomine [Bacteroides uniformis],5T99_B Crystal structure of BuGH2Awt in complex with Galactoisofagomine [Bacteroides uniformis] |
| 7CWD_A | 3.94e-68 | 16 | 354 | 461 | 799 | ChainA, beta-glalactosidase [Niallia circulans],7CWI_A Chain A, beta-galactosidase [Niallia circulans] |
| 4YPJ_A | 1.36e-66 | 2 | 346 | 449 | 797 | ChainA, Beta galactosidase [Niallia circulans],4YPJ_B Chain B, Beta galactosidase [Niallia circulans] |
| 4CU6_A | 3.88e-57 | 13 | 354 | 478 | 842 | Unravellingthe multiple functions of the architecturally intricate Streptococcus pneumoniae beta-galactosidase, BgaA [Streptococcus pneumoniae TIGR4],4CU7_A Unravelling the multiple functions of the architecturally intricate Streptococcus pneumoniae beta-galactosidase, BgaA [Streptococcus pneumoniae TIGR4],4CU8_A Unravelling the multiple functions of the architecturally intricate Streptococcus pneumoniae beta-galactosidase, BgaA [Streptococcus pneumoniae TIGR4] |
| 4CUC_A | 1.01e-56 | 13 | 354 | 478 | 842 | Unravellingthe multiple functions of the architecturally intricate Streptococcus pneumoniae beta-galactosidase, BgaA. [Streptococcus pneumoniae TIGR4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7LXS9 | 1.63e-64 | 13 | 354 | 513 | 846 | Beta-galactosidase BoGH2A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02645 PE=1 SV=1 |
| T2KM09 | 7.93e-26 | 77 | 307 | 547 | 770 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22050 PE=2 SV=2 |
| T2KN75 | 2.67e-15 | 72 | 354 | 523 | 793 | Beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22060 PE=1 SV=1 |
| T2KPJ7 | 1.94e-09 | 23 | 237 | 504 | 711 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
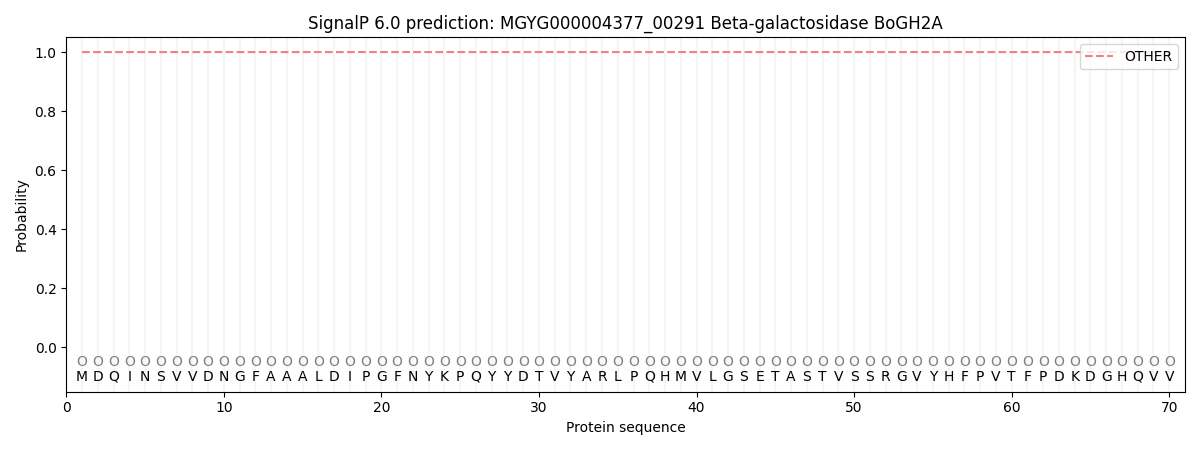
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000058 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
