You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004636_01639
You are here: Home > Sequence: MGYG000004636_01639
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paramuribaculum sp009775605 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; Paramuribaculum; Paramuribaculum sp009775605 | |||||||||||
| CAZyme ID | MGYG000004636_01639 | |||||||||||
| CAZy Family | GT51 | |||||||||||
| CAZyme Description | Monofunctional biosynthetic peptidoglycan transglycosylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3021; End: 3704 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT51 | 53 | 214 | 1.7e-45 | 0.903954802259887 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK00056 | mtgA | 1.17e-78 | 9 | 227 | 11 | 231 | monofunctional biosynthetic peptidoglycan transglycosylase; Provisional |
| TIGR02070 | mono_pep_trsgly | 5.05e-64 | 9 | 225 | 10 | 224 | monofunctional biosynthetic peptidoglycan transglycosylase. This family is one of the transglycosylases involved in the late stages of peptidoglycan biosynthesis. Members tend to be small, about 240 amino acids in length, and consist almost entirely of a domain described by pfam00912 for transglycosylases. Species with this protein will have several other transglycosylases as well. All species with this protein are Proteobacteria that produce murein (peptidoglycan). [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan] |
| pfam00912 | Transgly | 9.95e-53 | 52 | 224 | 9 | 173 | Transglycosylase. The penicillin-binding proteins are bifunctional proteins consisting of transglycosylase and transpeptidase in the N- and C-terminus respectively. The transglycosylase domain catalyzes the polymerization of murein glycan chains. |
| COG0744 | MrcB | 2.58e-50 | 8 | 215 | 23 | 235 | Membrane carboxypeptidase (penicillin-binding protein) [Cell wall/membrane/envelope biogenesis]. |
| COG5009 | MrcA | 2.15e-29 | 3 | 204 | 6 | 219 | Membrane carboxypeptidase/penicillin-binding protein [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AGY53411.1 | 1.16e-59 | 4 | 227 | 11 | 233 |
| AVR44737.1 | 1.15e-58 | 1 | 215 | 1 | 210 |
| BAR49817.1 | 2.24e-58 | 21 | 215 | 22 | 212 |
| AEW20603.1 | 2.24e-58 | 21 | 215 | 22 | 212 |
| BAR52639.1 | 2.24e-58 | 21 | 215 | 22 | 212 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2OQO_A | 1.27e-25 | 52 | 213 | 16 | 177 | Crystalstructure of a peptidoglycan glycosyltransferase from a class A PBP: insight into bacterial cell wall synthesis [Aquifex aeolicus VF5],3D3H_A Crystal structure of a complex of the peptidoglycan glycosyltransferase domain from Aquifex aeolicus and neryl moenomycin A [Aquifex aeolicus],3NB7_A Crystal structure of Aquifex Aeolicus Peptidoglycan Glycosyltransferase in complex with Decarboxylated Neryl Moenomycin [Aquifex aeolicus] |
| 3NB6_A | 2.50e-25 | 52 | 213 | 16 | 177 | Crystalstructure of Aquifex aeolicus peptidoglycan glycosyltransferase in complex with Methylphosphoryl Neryl Moenomycin [Aquifex aeolicus] |
| 4OON_A | 1.48e-18 | 37 | 201 | 20 | 184 | Crystalstructure of PBP1a in complex with compound 17 ((4Z,8S,11E,14S)-5-(2-amino-1,3-thiazol-4-yl)-14-(5,6-dihydroxy-1,3-dioxo-1,3-dihydro-2H-isoindol-2-yl)-8-formyl-2-methyl-6-oxo-3,10-dioxa-4,7,11-triazatetradeca-4,11-diene-2,12,14-tricarboxylic acid) [Pseudomonas aeruginosa PAO1] |
| 5U2G_A | 2.32e-17 | 59 | 215 | 43 | 199 | 2.6Angstrom Resolution Crystal Structure of Penicillin-Binding Protein 1A from Haemophilus influenzae [Haemophilus influenzae Rd KW20],5U2G_B 2.6 Angstrom Resolution Crystal Structure of Penicillin-Binding Protein 1A from Haemophilus influenzae [Haemophilus influenzae Rd KW20] |
| 3UDF_A | 6.34e-16 | 59 | 201 | 40 | 184 | ChainA, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDF_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDI_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDI_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDX_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDX_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE0_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE0_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE1_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE1_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q3V7R3 | 9.88e-54 | 17 | 220 | 19 | 222 | Biosynthetic peptidoglycan transglycosylase OS=Bdellovibrio bacteriovorus (strain ATCC 15356 / DSM 50701 / NCIMB 9529 / HD100) OX=264462 GN=mtgA PE=3 SV=1 |
| Q11VI0 | 7.52e-52 | 17 | 227 | 27 | 239 | Biosynthetic peptidoglycan transglycosylase OS=Cytophaga hutchinsonii (strain ATCC 33406 / DSM 1761 / CIP 103989 / NBRC 15051 / NCIMB 9469 / D465) OX=269798 GN=mtgA PE=3 SV=1 |
| B8GYX9 | 4.97e-49 | 18 | 218 | 21 | 215 | Biosynthetic peptidoglycan transglycosylase OS=Caulobacter vibrioides (strain NA1000 / CB15N) OX=565050 GN=mtgA PE=3 SV=1 |
| Q9ABA6 | 4.97e-49 | 18 | 218 | 21 | 215 | Biosynthetic peptidoglycan transglycosylase OS=Caulobacter vibrioides (strain ATCC 19089 / CB15) OX=190650 GN=mtgA PE=3 SV=1 |
| Q9CNV0 | 1.15e-48 | 15 | 215 | 29 | 230 | Biosynthetic peptidoglycan transglycosylase OS=Pasteurella multocida (strain Pm70) OX=272843 GN=mtgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
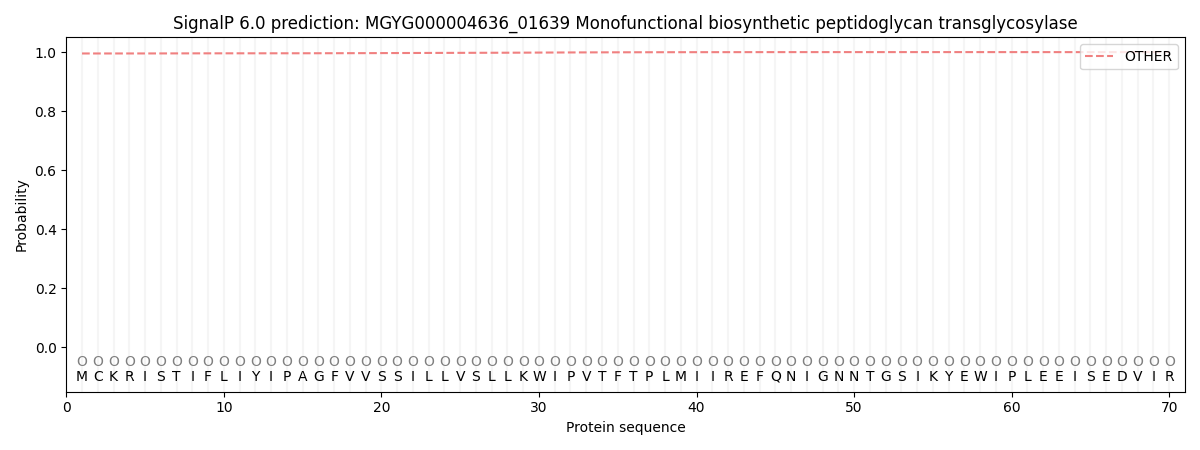
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.995633 | 0.004170 | 0.000048 | 0.000026 | 0.000014 | 0.000133 |

