

| Basic Information | |
|---|---|
| Species | Ricinus communis |
| Cazyme ID | 27504.m000617 |
| Family | CBM45 |
| Protein Properties | Length: 1469 Molecular Weight: 164321 Isoelectric Point: 7.1149 |
| Chromosome | Chromosome/Scaffold: 27504 Start: 59960 End: 71803 |
| Description | Pyruvate phosphate dikinase, PEP/pyruvate binding domain |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM45 | 131 | 206 | 4.7e-21 |
| LHWGGIRDRKEKWILPSRCPDGTKNYKNRALRSPFVKSGSSSYLKIEIDDPAIQALEFLVLDEGQNKWFKYKGQNF | |||
| CBM45 | 462 | 541 | 2.6e-21 |
| LHWALSRNSREWSAPPSGVLPPGSVTLSEAAETQLTNVSSAELPYQVQSFELEIEEDNFVGMPFVLLSNGNWIKNKGSDF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1469 Download |
| MSNSISHNLL QQSLVRHSVV LEHRNKLNSS SSSSAAASGI ASLSAPQIRR SSISSSFYGN 60 RLKISKSKLA IGTPRPATIT PRAVLAMDPA SELVGKFKLD GNSELQVSVS NAGSITQVNF 120 QISYGSDSLL LHWGGIRDRK EKWILPSRCP DGTKNYKNRA LRSPFVKSGS SSYLKIEIDD 180 PAIQALEFLV LDEGQNKWFK YKGQNFHVKL PEREKVMIQN VSVPEELVQV QAYLRWERKG 240 KQIYTPEQEK EEYDAARVEL LEELARGTSV EDLRTRLTNR NDRHEIKEPP VAETKTKIPD 300 DLVQIQSYIR WEKAGKPSYS PEQQLREFEE ARQDLQREVK RGVSLDEIRK KIAKGEIQSK 360 VSKQLQKQKY VSSEKIQRKR RDLAQLITKY AATPVEEPVS SEPKALKAIE LFAKAKEEQV 420 GGAVLNKKMF KLADGELLVL VTKPPGKTKI YVATDFREPV TLHWALSRNS REWSAPPSGV 480 LPPGSVTLSE AAETQLTNVS SAELPYQVQS FELEIEEDNF VGMPFVLLSN GNWIKNKGSD 540 FYIEFSGGPK QVQKDAGNGR GTAKALLDKI AEMESEAQKS FMHRFNIAAD LMEQAKDSGE 600 LGLAGILVWM RFMATRQLIW NKNYNVKPRE ISKAQDRLTD LLQNIYTSQP QYREILRMIM 660 STVGRGGEGD VGQRIRDEIL VIQRNNDCKG GMMEEWHQKL HNNTSPDDVV ICQALIDYIS 720 SGFDISMYWK SLNENGITKE RLLSYDRAIH SEPNFRRDQK DGLLRDLGNY MRTLKAVHSG 780 ADLESAIANC MGYRAEGQGF MVGVQINPIS GLPSGFPELL QFVLEHVEDK NVEALLEGLL 840 EARQELRPLL FKSHDRLKDL LFLDIALDST VRTVIERGYE ELNNAGQEKI MYFITLVLEN 900 LALSSDDNED LIYCMKGWNH ALSMSKSKSD QWALYAKSVL DRTRLALSSK AEWYQQVLQP 960 SAEYLGSLLG VDQWAVNIFT EEIIRAGSAA SLSSLLNRLD PILRKTANLG SWQVISPVEV 1020 AGYVVVVDEL LTVQNKSYGR PTILVARRVK GEEEIPDGTV AVLTPDMPDV LSHVSVRARN 1080 GKVCFATCFD HNILEKLQAH EGKLLQLKPT SADIVYNEIS EGELADSSST NMKEVGSSPI 1140 KLVKKQFSGR YAISSDEFTS EMVGAKSRNI SHLKGKVPSW IGIPTSVALP FGVFEKVLSD 1200 GSNKEVAKKL ELLKKKLGEG DFSVLGKIRE TVLGLAAPQQ LVQELKTSMQ SSGMPWPGDE 1260 GEQRWQQAWM AIKKVWASKW NERAYFSTRK VKLDHDYLCM AVLVQEIINA DYAFVIHTTN 1320 PSSGDSSEIY AEVVRGLGET LVGAYPGRAL SFVCKKQDLN SPQVLGYPSK PIGLFIRRSI 1380 IFRSDSNGED LEGYAGAGLY DSVPMDEEEK VVIDYSSDPL IMDGNFRQSI LSSIARAGSA 1440 IEELHGSAQD IEGVIRDGKL YVVQTRPQM |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
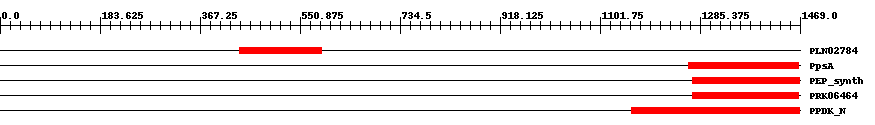 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02784 | PLN02784 | 1.0e-7 | 439 | 591 | 158 | + alpha-amylase | ||
| COG0574 | PpsA | 1.0e-9 | 1264 | 1467 | 207 | + Phosphoenolpyruvate synthase/pyruvate phosphate dikinase [Carbohydrate transport and metabolism] | ||
| TIGR01418 | PEP_synth | 4.0e-12 | 1271 | 1468 | 205 | + phosphoenolpyruvate synthase. Also called pyruvate,water dikinase and PEP synthase. The member from Methanococcus jannaschii contains a large intein. This enzyme generates phosphoenolpyruvate (PEP) from pyruvate, hydrolyzing ATP to AMP and releasing inorganic phosphate in the process. The enzyme shows extensive homology to other enzymes that use PEP as substrate or product. This enzyme may provide PEP for gluconeogenesis, for PTS-type carbohydrate transport systems, or for other processes [Energy metabolism, Glycolysis/gluconeogenesis]. | ||
| PRK06464 | PRK06464 | 1.0e-12 | 1271 | 1467 | 210 | + phosphoenolpyruvate synthase; Validated | ||
| pfam01326 | PPDK_N | 5.0e-27 | 1159 | 1468 | 337 | + Pyruvate phosphate dikinase, PEP/pyruvate binding domain. This enzyme catalyzes the reversible conversion of ATP to AMP, pyrophosphate and phosphoenolpyruvate (PEP). | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005524 | ATP binding |
| GO:0016301 | kinase activity |
| GO:0016310 | phosphorylation |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK11735.1 | 0 | 1 | 1469 | 1 | 1464 | starch associated protein R1 [Solanum tuberosum] |
| Swiss-Prot | Q8LPT9 | 0 | 1 | 1469 | 1 | 1475 | GWD1_CITRE RecName: Full=Alpha-glucan water dikinase, chloroplastic; AltName: Full=Starch-related protein R1; Flags: Precursor |
| RefSeq | XP_002270485.1 | 0 | 1 | 1469 | 1 | 1470 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002315679.1 | 0 | 12 | 1469 | 17 | 1477 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002527902.1 | 0 | 1 | 1469 | 1 | 1469 | alpha-glucan water dikinase, chloroplast precursor, putative [Ricinus communis] |
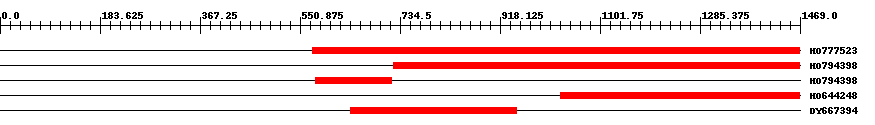
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO777523 | 897 | 574 | 1469 | 0 |
| HO794398 | 748 | 723 | 1469 | 0 |
| HO794398 | 141 | 579 | 719 | 0 |
| HO644248 | 443 | 1029 | 1469 | 0 |
| DY667394 | 306 | 644 | 949 | 0 |
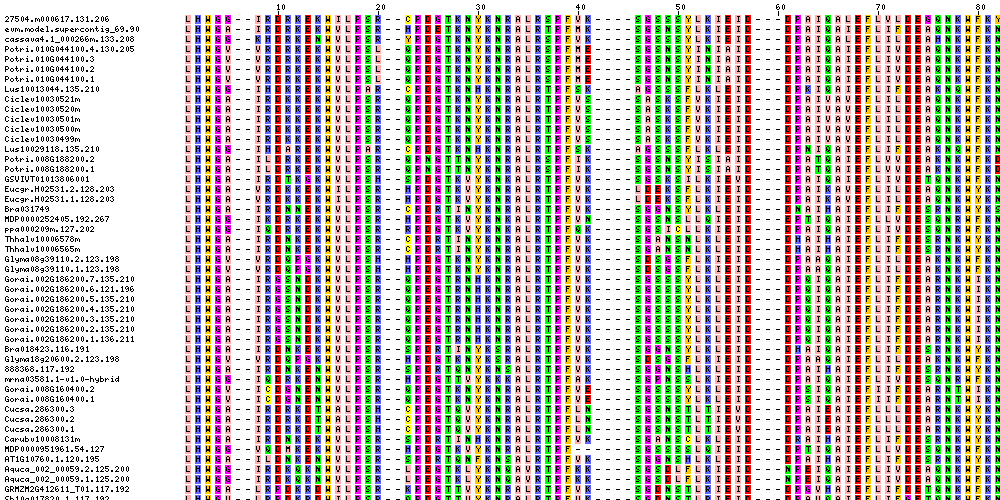
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |