

| Basic Information | |
|---|---|
| Species | Citrus clementina |
| Cazyme ID | Ciclev10010975m |
| Family | GT35 |
| Protein Properties | Length: 1002 Molecular Weight: 112838 Isoelectric Point: 4.564 |
| Chromosome | Chromosome/Scaffold: 6 Start: 25331673 End: 25338353 |
| Description | Glycosyl transferase, family 35 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT35 | 176 | 514 | 0 |
| ALSKLGQSLENVVSQEPDAALGNGGLGRLASCFLDSMATLNYPAWGYGLRYKYGLFKQRITKDGQEEVAEDWLELGNPWEIERNDVSYPVKFYGKIVPGS DGKSHWIGGEDIKAVAYDIPIPGYKTKTTINLRLWSTMVPSEDFDLSAFNAGDHTKAAEALTNAEKICYILYPGDESVEGKVLRLKQQYTLCSASLQDII ARFEKRSAANVNWEEFPEKVAVQMNDTHPTLCIPELIRILIDLKGLSWKEAWNITQRTVAYTNHTVLPEALEKWSFELMQKLLPRHMEIIEMIDEELVHT IVSEYGTADPDLLEKRLKEMRILENVDLPATFADLFVKT | |||
| GT35 | 585 | 995 | 0 |
| LEEEKEAEAVQEPPQLVRMANLCVVGSHAVNGVAEIHSEIVTNEVFNEFYKLWPEKFQNKTNGVTPRRWIRFCNPDLSSILTSWLGTEDWVTNTGKLAEL RKFADNEDLQSQFRAAKRNNKMKVVSFIKEKTGYSVSPDAMFDIQVKRIHEYKRQLMNILGIVYRYKKMKEMSAVERKAKFVPRVCIFGGKAFATYVQAK RIVKFITDVGATVNHDPEIGDLLKVIFVPDYNVSVAELLIPASELSQHISTAGMEASGTSNMKFAMNGCILIGTLDGANVEIRQEVGEENFFLFGARAHE IAGLRKERSEGKFVPDARFEEVKKFVKSGVFGSYNYDELMGSLEGNEGFGQADYFLVGKDFPSYLECQEKVDEAYCDQKRWTRMSIMNTAGSSKFSSDRT IQEYARDIWNI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1002 Download |
| MAVSQFSSMS TRPSEALVQC TSSLSRFIEF GSRNRTSKQK LLLIRTFNSR PPTTSFCIKC 60 VSSQPSPKTK DRVTEEDTSS SQNSSGPDTA SVASSIQYHA EFTPLFSPEK FEPPKAFFAT 120 AQSVRDSLII NWNSTYEYYE RLNVKQAYYL SMEFLQGRAL LNAIGNLGLT GAYAEALSKL 180 GQSLENVVSQ EPDAALGNGG LGRLASCFLD SMATLNYPAW GYGLRYKYGL FKQRITKDGQ 240 EEVAEDWLEL GNPWEIERND VSYPVKFYGK IVPGSDGKSH WIGGEDIKAV AYDIPIPGYK 300 TKTTINLRLW STMVPSEDFD LSAFNAGDHT KAAEALTNAE KICYILYPGD ESVEGKVLRL 360 KQQYTLCSAS LQDIIARFEK RSAANVNWEE FPEKVAVQMN DTHPTLCIPE LIRILIDLKG 420 LSWKEAWNIT QRTVAYTNHT VLPEALEKWS FELMQKLLPR HMEIIEMIDE ELVHTIVSEY 480 GTADPDLLEK RLKEMRILEN VDLPATFADL FVKTKESTDV VPDDELENCD EEGGPVDEEL 540 ESAQEDGVLE EESTDVVPDD ELENCDEEGG PVDEELESEQ EDDVLEEEKE AEAVQEPPQL 600 VRMANLCVVG SHAVNGVAEI HSEIVTNEVF NEFYKLWPEK FQNKTNGVTP RRWIRFCNPD 660 LSSILTSWLG TEDWVTNTGK LAELRKFADN EDLQSQFRAA KRNNKMKVVS FIKEKTGYSV 720 SPDAMFDIQV KRIHEYKRQL MNILGIVYRY KKMKEMSAVE RKAKFVPRVC IFGGKAFATY 780 VQAKRIVKFI TDVGATVNHD PEIGDLLKVI FVPDYNVSVA ELLIPASELS QHISTAGMEA 840 SGTSNMKFAM NGCILIGTLD GANVEIRQEV GEENFFLFGA RAHEIAGLRK ERSEGKFVPD 900 ARFEEVKKFV KSGVFGSYNY DELMGSLEGN EGFGQADYFL VGKDFPSYLE CQEKVDEAYC 960 DQKRWTRMSI MNTAGSSKFS SDRTIQEYAR DIWNIIPVEL P* |
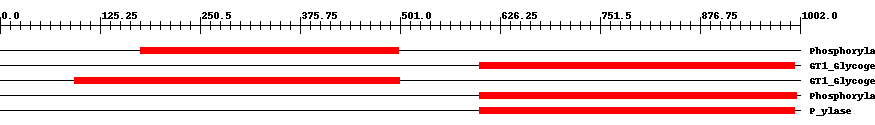
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00343 | Phosphorylase | 7.0e-141 | 176 | 499 | 324 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 601 | 995 | 404 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 93 | 500 | 412 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| pfam00343 | Phosphorylase | 0 | 601 | 997 | 402 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| TIGR02093 | P_ylase | 0 | 601 | 995 | 400 | + glycogen/starch/alpha-glucan phosphorylases. This family consists of phosphorylases. Members use phosphate to break alpha 1,4 linkages between pairs of glucose residues at the end of long glucose polymers, releasing alpha-D-glucose 1-phosphate. The nomenclature convention is to preface the name according to the natural substrate, as in glycogen phosphorylase, starch phosphorylase, maltodextrin phosphorylase, etc. Name differences among these substrates reflect differences in patterns of branching with alpha 1,6 linkages. Members include allosterically regulated and unregulated forms. A related family, TIGR02094, contains examples known to act well on particularly small alpha 1,4 glucans, as may be found after import from exogenous sources [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004645 | phosphorylase activity |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACJ11757.1 | 0 | 54 | 1001 | 2 | 935 | alpha-1,4 glucan phosphorylase [Gossypium hirsutum] |
| EMBL | CBI22291.1 | 0 | 23 | 1001 | 21 | 982 | unnamed protein product [Vitis vinifera] |
| Swiss-Prot | P53536 | 0 | 54 | 1000 | 58 | 1002 | PHSL_VICFA RecName: Full=Alpha-1,4 glucan phosphorylase L isozyme, chloroplastic/amyloplastic; AltName: Full=Starch phosphorylase L; Flags: Precursor |
| RefSeq | XP_002305367.1 | 0 | 53 | 1001 | 11 | 949 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002526085.1 | 0 | 19 | 1001 | 16 | 977 | glycogen phosphorylase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1qm5_B | 0 | 601 | 993 | 404 | 792 | A Chain A, Phosphorylase Recognition And Phosphorylysis Of Its Oligosaccharide Substrate: Answers To A Long Outstanding Question |
| PDB | 1qm5_B | 0 | 143 | 469 | 57 | 375 | A Chain A, Phosphorylase Recognition And Phosphorylysis Of Its Oligosaccharide Substrate: Answers To A Long Outstanding Question |
| PDB | 1qm5_A | 0 | 601 | 993 | 404 | 792 | A Chain A, Phosphorylase Recognition And Phosphorylysis Of Its Oligosaccharide Substrate: Answers To A Long Outstanding Question |
| PDB | 1qm5_A | 0 | 143 | 469 | 57 | 375 | A Chain A, Phosphorylase Recognition And Phosphorylysis Of Its Oligosaccharide Substrate: Answers To A Long Outstanding Question |
| PDB | 1e4o_B | 0 | 601 | 993 | 404 | 792 | A Chain A, Phosphorylase Recognition And Phosphorylysis Of Its Oligosaccharide Substrate: Answers To A Long Outstanding Question |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
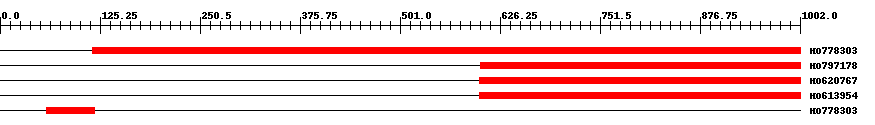 | ||||
| Hit | Length | Start | End | EValue |
| HO778303 | 887 | 116 | 1002 | 0 |
| HO797178 | 401 | 602 | 1002 | 0 |
| HO620767 | 403 | 600 | 1002 | 0 |
| HO613954 | 403 | 600 | 1002 | 0 |
| HO778303 | 63 | 58 | 118 | 0.0000004 |
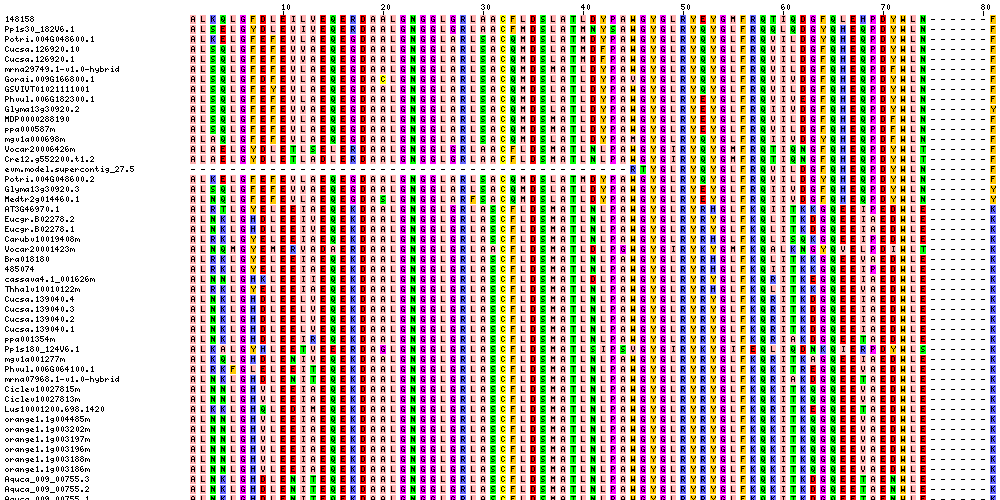
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |