

| Basic Information | |
|---|---|
| Species | Citrus clementina |
| Cazyme ID | Ciclev10031026m |
| Family | GH85 |
| Protein Properties | Length: 594 Molecular Weight: 66426.9 Isoelectric Point: 4.7492 |
| Chromosome | Chromosome/Scaffold: 4 Start: 1553215 End: 1558935 |
| Description | Glycosyl hydrolase family 85 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH85 | 1 | 68 | 1.6e-27 |
| MQGGYVDDKWVQGGTNADAYAIWHWHLMDVFVYFSHSLVTLPPPCWTNTAHRHGVKVLGTFITEGDEG | |||
| GH85 | 70 | 259 | 0 |
| LAVALGFDGWLINMEVELDVDQIPNLKEFVSHLTQTMHSSVPGSLAIWYDSVTIDGKLDWQDQLNEKNKPFFDICDGIFVNYTWKEDYPKLSAAVAGDRK FDVYMGIDVFGRNTFGGGKWNTNVALDVIKKDDVSAAIFAPGWIYETKQPPDFQTAQNHWWSLVEKSWGILQNYPKVLPFYSNFDQGHGL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 594 Download |
| MQGGYVDDKW VQGGTNADAY AIWHWHLMDV FVYFSHSLVT LPPPCWTNTA HRHGVKVLGT 60 FITEGDEGKL AVALGFDGWL INMEVELDVD QIPNLKEFVS HLTQTMHSSV PGSLAIWYDS 120 VTIDGKLDWQ DQLNEKNKPF FDICDGIFVN YTWKEDYPKL SAAVAGDRKF DVYMGIDVFG 180 RNTFGGGKWN TNVALDVIKK DDVSAAIFAP GWIYETKQPP DFQTAQNHWW SLVEKSWGIL 240 QNYPKVLPFY SNFDQGHGLH ISLEGEQMLD TPWNNISSQG FQPMLEFKDD PNPDTIQVLV 300 DFKEASYSGG GNVTFKGTLE ENASFIARLF QAELLLGNLP VYITYSVKSE ETSLLGLLLE 360 FSSARKERKS VLLASRRIDQ SSTKFSELIV PRQVKMLETT TEWATREARI IMGGYTITGI 420 SAVCYRPEPE NSRRTLESAS NVQDNASVHT PAEYFAILGD ISIKTSGQNS DFPLSSSWLV 480 EAQYVKFASD SQGTKTLSAK IIWKLKDGND SVFPQYNIYL GKPAKQAVGS LDGRVESTQE 540 YLGVARVESF YISDLIIPSN TDALKFIIQV CSVEGTSQNL DKSPFLLLDV KDK* 600 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG4724 | COG4724 | 4.0e-36 | 12 | 336 | 380 | + Endo-beta-N-acetylglucosaminidase D [Carbohydrate transport and metabolism] | ||
| pfam03644 | Glyco_hydro_85 | 4.0e-114 | 1 | 258 | 299 | + Glycosyl hydrolase family 85. Family of endo-beta-N-acetylglucosaminidases. These enzymes work on a broad spectrum of substrates. | ||
| cd06547 | GH85_ENGase | 8.0e-125 | 1 | 283 | 325 | + Endo-beta-N-acetylglucosaminidase (ENGase) hydrolyzes the N-N'-diacetylchitobiosyl core of N-glycosylproteins. The beta-1,4-glycosyl bond located between two N-acetylglucosamine residues is hydrolyzed such that N-acetylglucosamine 1 remains with the protein and N-acetylglucosamine 2 forms the reducing end of the released glycan. ENGase is a key enzyme in the processing of free oligosaccharides in the cytosol of eukaryotes. Oligosaccharides formed in the lumen of the endoplasmic reticulum are transported into the cytosol where they are catabolized by cytosolic ENGases and other enzymes, possibly to maximize the reutilization of the component sugars. ENGases have an eight-stranded alpha/beta barrel topology and are classified as a family 85 glycosyl hydrolase (GH85) domain. The GH85 ENGases are sequence-similar to the family 18 glycosyl hydrolases, also known as GH18 chitinases. An ENGase-like protein is also found in bacteria and is included in this alignment model. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005737 | cytoplasm |
| GO:0033925 | mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase activity |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI25424.1 | 0 | 28 | 593 | 1 | 592 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002273683.1 | 0 | 1 | 593 | 72 | 690 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002315137.1 | 0 | 1 | 591 | 76 | 697 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002520781.1 | 0 | 1 | 591 | 77 | 691 | endo beta n-acetylglucosaminidase, putative [Ricinus communis] |
| RefSeq | XP_002520784.1 | 0 | 1 | 593 | 73 | 687 | endo beta n-acetylglucosaminidase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3fhq_F | 8e-23 | 24 | 357 | 83 | 460 | A Chain A, Structure Of Endo-Beta-N-Acetylglucosaminidase A |
| PDB | 3fhq_D | 8e-23 | 24 | 357 | 83 | 460 | A Chain A, Structure Of Endo-Beta-N-Acetylglucosaminidase A |
| PDB | 3fhq_B | 8e-23 | 24 | 357 | 83 | 460 | A Chain A, Structure Of Endo-Beta-N-Acetylglucosaminidase A |
| PDB | 3fhq_A | 8e-23 | 24 | 357 | 83 | 460 | A Chain A, Structure Of Endo-Beta-N-Acetylglucosaminidase A |
| PDB | 3fha_D | 8e-23 | 24 | 357 | 83 | 460 | A Chain A, Structure Of Endo-Beta-N-Acetylglucosaminidase A |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
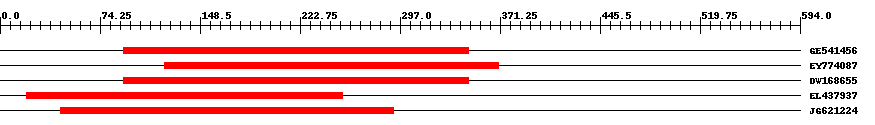 | ||||
| Hit | Length | Start | End | EValue |
| GE541456 | 257 | 92 | 348 | 0 |
| EY774087 | 249 | 122 | 370 | 0 |
| DW168655 | 257 | 92 | 348 | 0 |
| EL437937 | 257 | 20 | 254 | 0 |
| JG621224 | 271 | 45 | 292 | 0 |
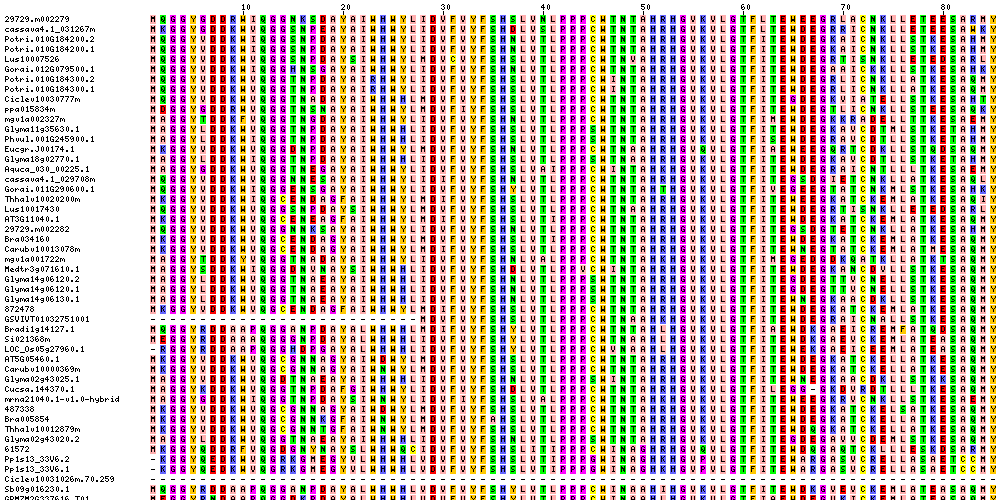
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |