

| Basic Information | |
|---|---|
| Species | Glycine max |
| Cazyme ID | Glyma13g27480.1 |
| Family | CBM53 |
| Protein Properties | Length: 1150 Molecular Weight: 130002 Isoelectric Point: 5.1454 |
| Chromosome | Chromosome/Scaffold: 13 Start: 30660060 End: 30670615 |
| Description | starch synthase 3 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
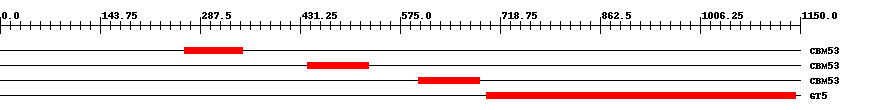 | |||
| Family | Start | End | Evalue |
| CBM53 | 265 | 348 | 1.8e-30 |
| QDIELFLNKNLSTLSEEPDILIMGAFNDWKWKSFSIRLNKLHLKGDWWSCQLYVPKEAYKVDFVFFNGQNVYDNNDQKDFCIPV | |||
| CBM53 | 442 | 529 | 2.5e-33 |
| IRLYYNRSSGPLANANEIWIHGGHNNWKYGLSIVERLVKSVLKGGEWWYADVVVPDQALVLDWVFADGPPKKAVVYDNNRKQDFHAIV | |||
| CBM53 | 601 | 689 | 2.3e-33 |
| GSTVTIFYNPSNTNLNGKPEVWFRCSFNRWSHRNGPLPPQRMLPAENGTHVKASFKVPLDAYMMDFVFSESEHGGVFDNKFGMDYHIPV | |||
| GT5 | 700 | 1143 | 0 |
| HIIHIAVEMAPIAKVGGLGDVVTSLSRAVQDLNHNVDIILPKYDCLNLSNVKDFDYHKSYSWGGTEIKVWHGKVEGLSVYFLEPQNGFFQVGCVYGRGND GERFGFFCHAALEFLLQNGFHPDIIHCHDWSSAPVAWLFKDNYAHYGLSKARVVFTIHNLEFGAHSIGKAMAYADKATTVSPTYSREIAGNPVIAPHLHK FHGIINGIDPDIWDPYNDKFIPVSYSSENVVEGKRASKETLQQRLSLKKADLPLVGIITRLTHQKGIHLIKHAIWRTLERGGQVVLLGSAPDPRIQNDFV NLANELHSAHHDRARLCLAYDEPLSHLIYAGADFILVPSIFEPCGLTQLTAMRYGSIPVVRKTGGLYDTVFDVDHDKDRAQAQGLEPNGFSFDGADTGGV DYALNRAISAWYEGRDWFNSLCKRVMEQDWSWNRPALDYLELYH | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1150 Download |
| MEMSMQLNCK TVFPYRGGYI CSPCKAGWGV SFVRASADFS RKRQQKKVSV ARTKGTSGKG 60 FVPSKKNTRM KKGDTLTSVV SEVSGGDKKQ TVEVNVDDTD KEGELEFSQE EKFEAVDRID 120 ENVGDVGDLS LLDETVGELS LLDESNQATI SVFDEDVEVL ESWKEEFPYN GGVGIVEDSS 180 EEGLLESAEI DENVKDTDTD GDITEEAVEE SSSADDDRIN EEAAGLLKLE LEANQRRQEI 240 ERIAEEKLSQ GIKLFVYPPV VKPDQDIELF LNKNLSTLSE EPDILIMGAF NDWKWKSFSI 300 RLNKLHLKGD WWSCQLYVPK EAYKVDFVFF NGQNVYDNND QKDFCIPVDG GMDALAFEDF 360 LLEEKRKELE ELARAQAERE RQAEEQRRIE ADRAAKEEDR ARAKAEIGKM RETLPQLLKN 420 AVKSVDNVWH IEPSEFKGKD LIRLYYNRSS GPLANANEIW IHGGHNNWKY GLSIVERLVK 480 SVLKGGEWWY ADVVVPDQAL VLDWVFADGP PKKAVVYDNN RKQDFHAIVP GAIPDEQYWV 540 EEEQLIYRKF QEERRLREDA IRAKAEKTAQ MKAETKERTL KGFLLSQKHI VFTDPLDVQA 600 GSTVTIFYNP SNTNLNGKPE VWFRCSFNRW SHRNGPLPPQ RMLPAENGTH VKASFKVPLD 660 AYMMDFVFSE SEHGGVFDNK FGMDYHIPVF GSIAKEPPLH IIHIAVEMAP IAKVGGLGDV 720 VTSLSRAVQD LNHNVDIILP KYDCLNLSNV KDFDYHKSYS WGGTEIKVWH GKVEGLSVYF 780 LEPQNGFFQV GCVYGRGNDG ERFGFFCHAA LEFLLQNGFH PDIIHCHDWS SAPVAWLFKD 840 NYAHYGLSKA RVVFTIHNLE FGAHSIGKAM AYADKATTVS PTYSREIAGN PVIAPHLHKF 900 HGIINGIDPD IWDPYNDKFI PVSYSSENVV EGKRASKETL QQRLSLKKAD LPLVGIITRL 960 THQKGIHLIK HAIWRTLERG GQVVLLGSAP DPRIQNDFVN LANELHSAHH DRARLCLAYD 1020 EPLSHLIYAG ADFILVPSIF EPCGLTQLTA MRYGSIPVVR KTGGLYDTVF DVDHDKDRAQ 1080 AQGLEPNGFS FDGADTGGVD YALNRAISAW YEGRDWFNSL CKRVMEQDWS WNRPALDYLE 1140 LYHAARKAE* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02316 | PLN02316 | 6.0e-6 | 554 | 693 | 143 | + synthase/transferase | ||
| TIGR02095 | glgA | 2.0e-158 | 699 | 1145 | 489 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| PRK00654 | glgA | 2.0e-169 | 699 | 1148 | 487 | + glycogen synthase; Provisional | ||
| PLN02316 | PLN02316 | 0 | 248 | 1147 | 900 | + synthase/transferase | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 0 | 700 | 1143 | 489 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| GO:2001070 | starch binding |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAF49176.1 | 0 | 1 | 1149 | 1 | 1165 | starch synthase III [Phaseolus vulgaris] |
| EMBL | CAB40374.1 | 0 | 1 | 1149 | 1 | 1147 | Starch synthase isoform SS III [Vigna unguiculata] |
| RefSeq | XP_002269011.1 | 0 | 45 | 1147 | 32 | 1140 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002305571.1 | 0 | 208 | 1147 | 63 | 1017 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002518476.1 | 0 | 48 | 1147 | 1 | 1058 | starch synthase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3d1j_A | 0 | 699 | 1143 | 1 | 474 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 3guh_A | 0 | 699 | 1143 | 1 | 474 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2r4u_A | 0 | 699 | 1143 | 1 | 474 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2r4t_A | 0 | 699 | 1143 | 1 | 474 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2qzs_A | 0 | 699 | 1143 | 1 | 474 | A Chain A, Crystal Structure Of Wild-Type E.Coli Gs In Complex With Adp And Glucose(Wtgsb) |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| starch biosynthesis | GLYCOGENSYN-RXN | EC-2.4.1.21 | starch synthase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
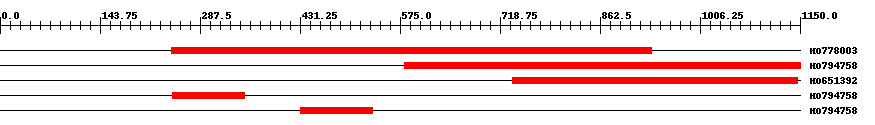 | ||||
| Hit | Length | Start | End | EValue |
| HO778003 | 690 | 247 | 936 | 0 |
| HO794758 | 570 | 581 | 1150 | 0 |
| HO651392 | 412 | 736 | 1147 | 0 |
| HO794758 | 109 | 248 | 352 | 0.004 |
| HO794758 | 107 | 432 | 536 | 1.9 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |