

| Basic Information | |
|---|---|
| Species | Glycine max |
| Cazyme ID | Glyma19g03611.1 |
| Family | GT1 |
| Protein Properties | Length: 430 Molecular Weight: 48876.1 Isoelectric Point: 6.5618 |
| Chromosome | Chromosome/Scaffold: 19 Start: 3650849 End: 3660216 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 66 | 202 | 1.3e-24 |
| NNLMVVLYVTFGSFTHFDQNQFHELALGLDLTNRPFLWVNKLEYPNEFLGTKGNIVGWAPQQKVLSHPAIACFATHCGWNSIMEGLSNGVLLLCWPYFAD QLYNKTHICDELKVGLGFEKDKNGLVSREEFKMKVDQ | |||
| GT1 | 284 | 406 | 7.4e-29 |
| MSWLDQQPHCSNTYVAFGHKGKIIGWAPQQKVLSHPAVACFISHCGWNSSTECLSNGVPFLCWPYFGDQPYNRKYICYELNVGLGLNSNENGLVSRWEIK KKLNQLLSDENIKSRSLKLKEKV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 430 Download |
| MPTMLEKLIE DIHLNGDNRI SLIVADLCIG WALNFGAKFG IFALVYNLPK LIDDGIIDSD 60 GVSLINNLMV VLYVTFGSFT HFDQNQFHEL ALGLDLTNRP FLWVNKLEYP NEFLGTKGNI 120 VGWAPQQKVL SHPAIACFAT HCGWNSIMEG LSNGVLLLCW PYFADQLYNK THICDELKVG 180 LGFEKDKNGL VSREEFKMKV DQFFNDENIK SSAHFLNYLV HHCTPALSLT EWWLSNTAYE 240 LEPWMLTLSP KLLPIGPLLR SYDNTNATTL RSLGQFWEED LSCMSWLDQQ PHCSNTYVAF 300 GHKGKIIGWA PQQKVLSHPA VACFISHCGW NSSTECLSNG VPFLCWPYFG DQPYNRKYIC 360 YELNVGLGLN SNENGLVSRW EIKKKLNQLL SDENIKSRSL KLKEKVTSNT TNRGQSLENF 420 NKFVKWLKE* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02448 | PLN02448 | 4.0e-25 | 234 | 425 | 244 | + UDP-glycosyltransferase family protein | ||
| PLN02562 | PLN02562 | 6.0e-28 | 250 | 428 | 225 | + UDP-glycosyltransferase | ||
| PLN02448 | PLN02448 | 3.0e-30 | 71 | 211 | 145 | + UDP-glycosyltransferase family protein | ||
| PLN02555 | PLN02555 | 3.0e-34 | 71 | 200 | 145 | + limonoid glucosyltransferase | ||
| PLN02555 | PLN02555 | 3.0e-36 | 234 | 428 | 257 | + limonoid glucosyltransferase | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACU24307.1 | 2e-18 | 1 | 72 | 92 | 175 | unknown [Glycine max] |
| GenBank | ACU24307.1 | 0 | 71 | 213 | 280 | 426 | unknown [Glycine max] |
| GenBank | ACU24307.1 | 0 | 158 | 429 | 136 | 456 | unknown [Glycine max] |
| GenBank | ACZ74670.1 | 0 | 71 | 210 | 286 | 429 | UDP-glucosyl transferase [Phaseolus vulgaris] |
| GenBank | ACZ74670.1 | 0 | 179 | 429 | 167 | 462 | UDP-glucosyl transferase [Phaseolus vulgaris] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2pq6_A | 1e-30 | 234 | 425 | 230 | 475 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2pq6_A | 2e-23 | 71 | 202 | 297 | 435 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2acv_B | 2e-22 | 71 | 209 | 278 | 428 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 2acv_B | 1e-18 | 250 | 424 | 239 | 458 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 2acv_A | 2e-22 | 71 | 209 | 278 | 428 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
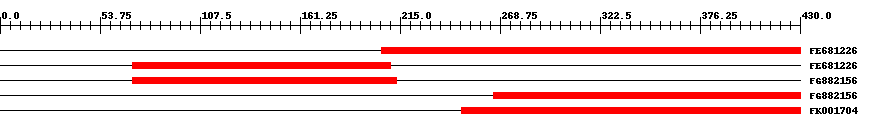 | ||||
| Hit | Length | Start | End | EValue |
| FE681226 | 270 | 205 | 430 | 0 |
| FE681226 | 144 | 71 | 210 | 0 |
| FG882156 | 147 | 71 | 213 | 0 |
| FG882156 | 208 | 265 | 430 | 0 |
| FK001704 | 229 | 248 | 430 | 0 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
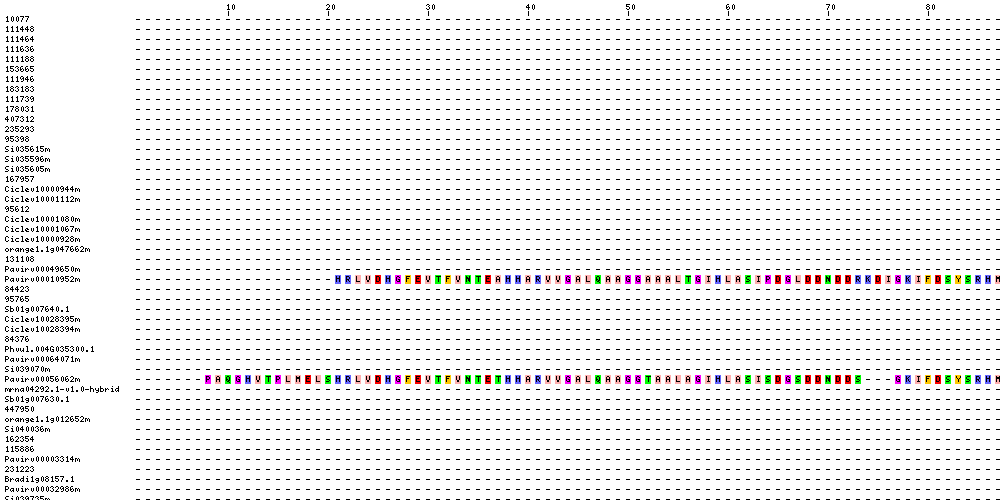 |