

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.010G239400.2 |
| Family | GT5 |
| Protein Properties | Length: 1014 Molecular Weight: 114965 Isoelectric Point: 5.6777 |
| Chromosome | Chromosome/Scaffold: 10 Start: 60896240 End: 60904968 |
| Description | starch synthase 4 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 555 | 786 | 0 |
| HVIHIAAEMAPVAKVGGLGDVVTGLGKALQKKGHLVEIVLPKYDCMQYDRIRDLRVLDATVYSYFDGKLFQNKVWTGTVEGLPVYFIEPHHPSKFFWRGQ YYGEQDDFKRFSFFSRAALELLLQVGKKPDIIHCHDWQTAFVAPLYWDLYFPKGLNSARICFTCHNFEYQGAAPASELASCGLDVQQLHRPDRMQDNSAH DRVNPIKGAIVFSNIVTTVSPTYAQEVRTAEY | |||
| GT5 | 784 | 1000 | 0 |
| AEYTANDLQGKAENKAAMRRHLRLSSADDSQPLVGCITRLVPQKGVHLIRHAIYRTLEMGGQFVLLGSSPVPHIQREFEGIANQFQDHEHIRLILKYDES LSRYIYAASDMFIIPSIFEPCGLTQMIAMRYGSVPIVRKTGGLNDSVFDVDDDTIPYQYRNGFTFATPDEQGLNGALDRAFNLYNNDSETWQQLVQKNMN IDFSWSSSASQYEELYA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1014 Download |
| MAARCSICFF SYGFNGLSEG NYGNDVDCWK KNVSLFLPSR RRLLPVCCKI QRRNLSSYNR 60 RQGKKPPFER IRPSAKLQPN SDDEFDPEHS VPNNGDMEPS VNREKTFEDD VDARGDAEHT 120 DEKNLGTQFA SAIETNRDVK HADEQITDSP AQSAVAKASA INGVGAELLS SVQPDDLIGM 180 IKNAERNILL LNQARVHALE DLHKILSEKE TLKGEINNLE KRLAEADAQI KFASQEKVHA 240 ELLEDQLENL QNELINRGGS GKSELELYEN RSKISNEGAL LARDGHIHSL SKEVDSLRTE 300 NLALKYDIQA LKSMLSNLKN TDKRIVTLEN ESSFLESSVK ELESKLSVSQ QESSNISTLK 360 TECKDLWAKV ENLQLLLDKA TKQADQAILV LQQNQDLRKK VDKLEESLEA ATIFKASSEK 420 TQQYNELMQQ KIKLLEERLQ KSDEEIYSYV QLYQESIKEF RDTLNSLKEE SKKRALDEPV 480 DDMPWEFWSC LLLTIDGWVL ENKILNSEAV PLREMVWKRD RRICDAYVIC KEKTEDEVIS 540 TFLQLISSQA SPGLHVIHIA AEMAPVAKVG GLGDVVTGLG KALQKKGHLV EIVLPKYDCM 600 QYDRIRDLRV LDATVYSYFD GKLFQNKVWT GTVEGLPVYF IEPHHPSKFF WRGQYYGEQD 660 DFKRFSFFSR AALELLLQVG KKPDIIHCHD WQTAFVAPLY WDLYFPKGLN SARICFTCHN 720 FEYQGAAPAS ELASCGLDVQ QLHRPDRMQD NSAHDRVNPI KGAIVFSNIV TTVSPTYAQE 780 VRTAEYTAND LQGKAENKAA MRRHLRLSSA DDSQPLVGCI TRLVPQKGVH LIRHAIYRTL 840 EMGGQFVLLG SSPVPHIQRE FEGIANQFQD HEHIRLILKY DESLSRYIYA ASDMFIIPSI 900 FEPCGLTQMI AMRYGSVPIV RKTGGLNDSV FDVDDDTIPY QYRNGFTFAT PDEQGLNGAL 960 DRAFNLYNND SETWQQLVQK NMNIDFSWSS SASQYEELYA KSVARARAAT SRT* 1020 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02316 | PLN02316 | 3.0e-126 | 552 | 999 | 487 | + synthase/transferase | ||
| PRK00654 | glgA | 2.0e-159 | 554 | 999 | 488 | + glycogen synthase; Provisional | ||
| TIGR02095 | glgA | 5.0e-170 | 554 | 1002 | 491 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 1.0e-170 | 555 | 1001 | 491 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| PLN02939 | PLN02939 | 0 | 46 | 1003 | 997 | + transferase, transferring glycosyl groups | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACT09058.1 | 0 | 32 | 1010 | 25 | 999 | starch synthase IV precursor [Solanum lycopersicum] |
| EMBL | CAA16796.1 | 0 | 6 | 1013 | 7 | 1071 | starch synthase-like protein [Arabidopsis thaliana] |
| RefSeq | NP_193558.3 | 0 | 6 | 1013 | 7 | 1040 | ATSS4; transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | XP_002274716.1 | 0 | 1 | 1010 | 1 | 1009 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002298514.1 | 0 | 156 | 1012 | 14 | 887 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3d1j_A | 0 | 554 | 1001 | 1 | 475 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 3guh_A | 0 | 554 | 1001 | 1 | 475 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2r4u_A | 0 | 554 | 1001 | 1 | 475 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2r4t_A | 0 | 554 | 1001 | 1 | 475 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2qzs_A | 0 | 554 | 1001 | 1 | 475 | A Chain A, Crystal Structure Of Wild-Type E.Coli Gs In Complex With Adp And Glucose(Wtgsb) |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| GO851132 | 332 | 629 | 922 | 0 |
| EL448877 | 304 | 482 | 785 | 0 |
| CF828776 | 285 | 680 | 925 | 0 |
| GW865616 | 298 | 660 | 918 | 0 |
| GO859521 | 226 | 777 | 1002 | 0 |
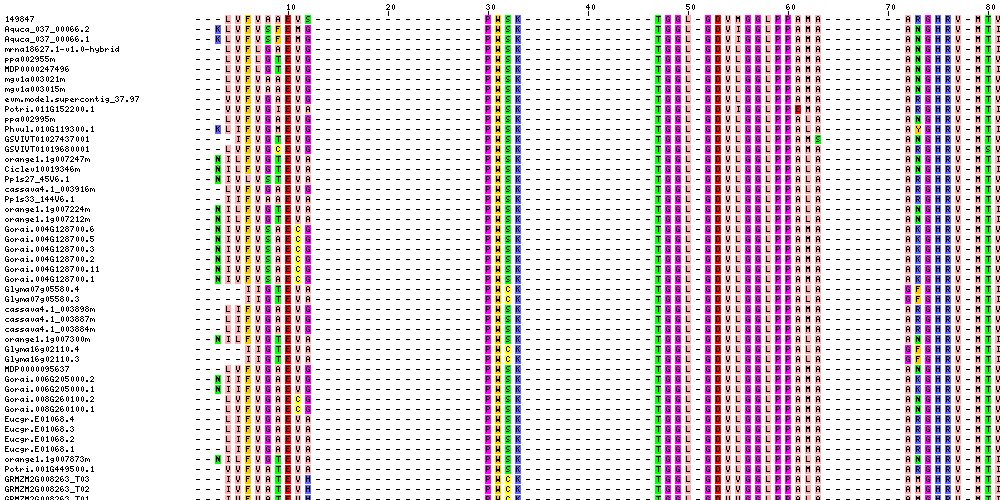
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |