

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.011G077500.1 |
| Family | GT4 |
| Protein Properties | Length: 698 Molecular Weight: 78240.2 Isoelectric Point: 10.6754 |
| Chromosome | Chromosome/Scaffold: 11 Start: 7648877 End: 7654557 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT4 | 546 | 666 | 3.6e-23 |
| VGSKSNKIPYVKEILSFLSQHAKLSESVLWTPATTRVASLYSAADVYVMNSQGLGETFGRVTVEAMAFGLPVLGTDGGGTKEIVEHNVTGLLHPMGHPGT RVLAENLRFLLKNLNARKQMG | |||
| Full Sequence |
|---|
| Protein Sequence Length: 698 Download |
| MEERLSKGPS SLRQGSLKSS LSGRSTPKGS PTYRRLNSSR TPRREARSGA GGTQWFRSNR 60 VVYWLLLITL WAYLGFYVQS RWAHGHKKEE FLGFNGDPRD KLVDAEQNTR RDLLTDDSLV 120 AVNNITNKTQ VHVDRKIDVI LAKKGNGFTS RKKRSKRRRR NLPKVRDKLK AKTNTESGDA 180 EGQELEILQK NSTFGLLVGP FGSLEDRVLE WSPEKRSGTC DRKGDFARLV WSRRLVLVFH 240 ELSMTGAPIS MMELATEFLS CGATVSAVVL SKKGGLMSEL ARRRIKVIED RADLSFKTAM 300 KADLVIAGSA VCASWIDQYI AHFPAGGSQI AWWIMENRRE YFDRSKLVLH RVKMLIFLSE 360 LQSKQWLTWC QEENIKLRSQ PALVPLAVND ELAFVAGFPC SLNTPSASSV KMLEKRQLLR 420 DAARKEMGLT DNDMLVISLS SINAGKGQLF LLESADLAMN EDPLQTGSEV KKSLDIRQDQ 480 PSLSVKHHLR GLHQKSRNLD VSSTNLRLFT SVNTTNAVSI NGTHRRKMYD SKGAQEQALK 540 ILIGSVGSKS NKIPYVKEIL SFLSQHAKLS ESVLWTPATT RVASLYSAAD VYVMNSQGLG 600 ETFGRVTVEA MAFGLPVLGT DGGGTKEIVE HNVTGLLHPM GHPGTRVLAE NLRFLLKNLN 660 ARKQMGMEGR KMVERKYLKR HMYKRFVEVL TKCMRSK* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
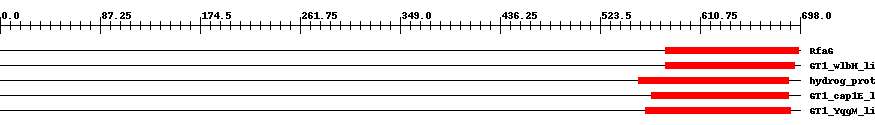 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0438 | RfaG | 6.0e-20 | 581 | 697 | 117 | + Glycosyltransferase [Cell envelope biogenesis, outer membrane] | ||
| cd03798 | GT1_wlbH_like | 1.0e-20 | 581 | 693 | 113 | + This family is most closely related to the GT1 family of glycosyltransferases. wlbH in Bordetella parapertussis has been shown to be required for the biosynthesis of a trisaccharide that, when attached to the B. pertussis lipopolysaccharide (LPS) core (band B), generates band A LPS. | ||
| TIGR00072 | hydrog_prot | 3.0e-21 | 557 | 688 | 133 | + hydrogenase maturation protease. HycI and HoxM are well-characterized as responsible for C-terminal protease activity on their respective hydrogenase large chains. A large number of homologous proteins appear responsible for the maturation of various forms of hydrogenase. | ||
| cd03808 | GT1_cap1E_like | 2.0e-23 | 568 | 688 | 121 | + This family is most closely related to the GT1 family of glycosyltransferases. cap1E in Streptococcus pneumoniae is required for the synthesis of type 1 capsular polysaccharides. | ||
| cd03801 | GT1_YqgM_like | 2.0e-27 | 563 | 690 | 130 | + This family is most closely related to the GT1 family of glycosyltransferases and named after YqgM in Bacillus licheniformis about which little is known. Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. This group of glycosyltransferases is most closely related to the previously defined glycosyltransferase family 1 (GT1). The members of this family may transfer UDP, ADP, GDP, or CMP linked sugars. The diverse enzymatic activities among members of this family reflect a wide range of biological functions. The protein structure available for this family has the GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. The members of this family are found mainly in certain bacteria and archaea. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAN71826.1 | 0 | 10 | 697 | 18 | 734 | hypothetical protein [Vitis vinifera] |
| EMBL | CBI36173.1 | 0 | 10 | 697 | 18 | 683 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002284822.1 | 0 | 10 | 697 | 7 | 691 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002298139.1 | 0 | 5 | 697 | 15 | 681 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002528176.1 | 0 | 15 | 697 | 24 | 686 | glycosyltransferase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2jjm_L | 0.00000004 | 563 | 695 | 259 | 387 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2jjm_K | 0.00000004 | 563 | 695 | 259 | 387 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2jjm_J | 0.00000004 | 563 | 695 | 259 | 387 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2jjm_I | 0.00000004 | 563 | 695 | 259 | 387 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2jjm_H | 0.00000004 | 563 | 695 | 259 | 387 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |