

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_10332686g0010 |
| Family | PL1 |
| Protein Properties | Length: 251 Molecular Weight: 27598.7 Isoelectric Point: 9.0839 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL1 | 134 | 249 | 7.7e-27 |
| AVIQTEPLWIIFRNDMVIQLKEELIMNSFKTIDGRGANVHIGNGPCITVQNVTNIIIHGIHIHDCKRAGNAMVKDSPQHTGWRTVSDGDGISIFTGRHIW VDHCSMSNCEDGLIDA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 251 Download |
| MLLIGILCCV HGGAAKPFGS KRGEKFMGRR KVAPGNITRV MFPVAKHPGN ATQIVRDRST 60 INSTRRGLLS CATGNPIDDC WRCDPNWERN RKRLADCGIG FGRDAIGGKN GKVYVVTDPR 120 DDPVNPSNGT LRHAVIQTEP LWIIFRNDMV IQLKEELIMN SFKTIDGRGA NVHIGNGPCI 180 TVQNVTNIII HGIHIHDCKR AGNAMVKDSP QHTGWRTVSD GDGISIFTGR HIWVDHCSMS 240 NCEDGLIDAV K |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG3866 | PelB | 2.0e-6 | 161 | 251 | 100 | + Pectate lyase [Carbohydrate transport and metabolism] | ||
| pfam00544 | Pec_lyase_C | 2.0e-23 | 154 | 251 | 121 | + Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. | ||
| smart00656 | Amb_all | 3.0e-26 | 148 | 251 | 115 | + Amb_all domain. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABR17766.1 | 0 | 1 | 250 | 10 | 254 | unknown [Picea sitchensis] |
| GenBank | ABR17909.1 | 0 | 1 | 251 | 10 | 258 | unknown [Picea sitchensis] |
| GenBank | ABR18471.1 | 0 | 1 | 250 | 10 | 269 | unknown [Picea sitchensis] |
| GenBank | ABV32549.1 | 0 | 43 | 250 | 31 | 241 | pectase lyase [Prunus persica] |
| DDBJ | BAE48664.1 | 0 | 43 | 250 | 31 | 241 | Pectate lyase [Prunus mume] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1pxz_B | 0 | 75 | 248 | 2 | 177 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1pxz_A | 0 | 75 | 248 | 2 | 177 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 3zsc_A | 0.0005 | 143 | 251 | 50 | 146 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| Signal Peptide | ||||
|---|---|---|---|---|
 | ||||
| Cleavage Site | ||||
| 0 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR491918 | 254 | 1 | 250 | 0 |
| GW772379 | 215 | 1 | 210 | 0 |
| FG140358 | 194 | 60 | 250 | 0 |
| EB427443 | 194 | 60 | 250 | 0 |
| ES873452 | 207 | 48 | 250 | 0 |
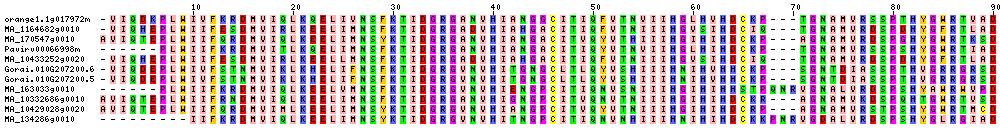
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |