

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_1164682g0010 |
| Family | PL1 |
| Protein Properties | Length: 266 Molecular Weight: 29649.4 Isoelectric Point: 6.3803 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL1 | 119 | 262 | 4.8e-38 |
| VIQHEPLWIIFESDMVIRLKEELIMNSFKTIDGRGADVHIAHGACITIQFVTNIIIHGVSIHDCIQTGNAMVRDSPDHYGFRTLADGDGISIFGGRFIWI DHCSFSSCTHGLIDAIMGSTAITISNNYFTEHNMVMLLGHSDSY | |||
| Full Sequence |
|---|
| Protein Sequence Length: 266 Download |
| MGFLRGRPTI QAAQQQTQHH KHNSSFPVES PEEVVKMVQK NINDSRRQLT YLSCGTGNPI 60 DDCWRCDPNW QMNRQRLADC AIGFGRNAIG GKNGQYYVVT DSSDNDAVNP TPGTLRHAVI 120 QHEPLWIIFE SDMVIRLKEE LIMNSFKTID GRGADVHIAH GACITIQFVT NIIIHGVSIH 180 DCIQTGNAMV RDSPDHYGFR TLADGDGISI FGGRFIWIDH CSFSSCTHGL IDAIMGSTAI 240 TISNNYFTEH NMVMLLGHSD SYVQDV |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG3866 | PelB | 5.0e-12 | 145 | 261 | 126 | + Pectate lyase [Carbohydrate transport and metabolism] | ||
| pfam00544 | Pec_lyase_C | 2.0e-32 | 138 | 265 | 150 | + Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. | ||
| smart00656 | Amb_all | 9.0e-40 | 132 | 265 | 145 | + Amb_all domain. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABR18471.1 | 0 | 1 | 266 | 39 | 301 | unknown [Picea sitchensis] |
| EMBL | CAN76500.1 | 0 | 24 | 265 | 21 | 262 | hypothetical protein [Vitis vinifera] |
| EMBL | CBI14899.1 | 0 | 24 | 265 | 21 | 262 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002275781.1 | 0 | 24 | 265 | 76 | 317 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002285340.1 | 0 | 24 | 265 | 21 | 262 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1pxz_B | 0 | 58 | 265 | 2 | 210 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1pxz_A | 0 | 58 | 265 | 2 | 210 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1vbl_A | 0.000008 | 141 | 265 | 128 | 266 | A Chain A, Structure Of The Thermostable Pectate Lyase Pl 47 |
| PDB | 3zsc_A | 0.00001 | 127 | 261 | 50 | 172 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| PDB | 2o1d_A | 0.00002 | 206 | 265 | 184 | 260 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EX393983 | 266 | 1 | 266 | 0 |
| DR479712 | 234 | 1 | 234 | 0 |
| CV531518 | 244 | 24 | 265 | 0 |
| DT546935 | 242 | 24 | 265 | 0 |
| DT557086 | 242 | 24 | 265 | 0 |
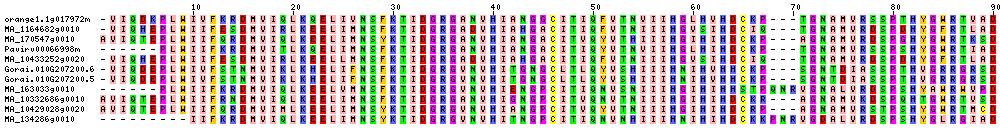
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |