

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_143862g0080 |
| Family | GH110 |
| Protein Properties | Length: 601 Molecular Weight: 65246.2 Isoelectric Point: 5.951 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH110 | 32 | 564 | 1.4013e-45 |
| SVQRGIDQAYRGGLHKVVVPPGVYHLTTESSSGSHLHLAGTKDFEIDATGATFILSTPRKGGVDFHGCQDVTLKGATLVRDPLPFSQGRIVALGADYQQI DVQVDKGYPADFDDKQSYAKLSLNSYDGEHNRRWEDDLWSARDVERLGPDTFRFHAGRCVGADSTWKMGSAVAWRGWGGADIAVTGCRGMRIDGVTIKNG AGFCVAESGGEGGNYYNYTVTYGPRPEGATAEPILASNADAFHSKSVRKGPTLEDCHFEGMNDDGIPIHGQYALIEQVEGNQFIVETKNYPFCQAGDHLR FLDERGALAGEARVVSVVDVPVFTPPAPAPRDLRLFQDLTKHQYTRITLDTAAAAKFGWLVANADANGDGFVVRRCTIRSHRARGMLIKASDGLIEDCTV EGSSIGGIVIAPEMDFWDESDFARNIVVRHNTIRDCDYWHQEGTSQAGAMTVDAFVAKRFVPLPGGHRHLLIEDNTFEENNGTNLVVSSAQDVIIRRNRF VRPMQKENGRGKARGVNPQALIWLTECSGIQLL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 601 Download |
| MVFSLVAVMW QSWNLLVQGA PPIVISPADP ASVQRGIDQA YRGGLHKVVV PPGVYHLTTE 60 SSSGSHLHLA GTKDFEIDAT GATFILSTPR KGGVDFHGCQ DVTLKGATLV RDPLPFSQGR 120 IVALGADYQQ IDVQVDKGYP ADFDDKQSYA KLSLNSYDGE HNRRWEDDLW SARDVERLGP 180 DTFRFHAGRC VGADSTWKMG SAVAWRGWGG ADIAVTGCRG MRIDGVTIKN GAGFCVAESG 240 GEGGNYYNYT VTYGPRPEGA TAEPILASNA DAFHSKSVRK GPTLEDCHFE GMNDDGIPIH 300 GQYALIEQVE GNQFIVETKN YPFCQAGDHL RFLDERGALA GEARVVSVVD VPVFTPPAPA 360 PRDLRLFQDL TKHQYTRITL DTAAAAKFGW LVANADANGD GFVVRRCTIR SHRARGMLIK 420 ASDGLIEDCT VEGSSIGGIV IAPEMDFWDE SDFARNIVVR HNTIRDCDYW HQEGTSQAGA 480 MTVDAFVAKR FVPLPGGHRH LLIEDNTFEE NNGTNLVVSS AQDVIIRRNR FVRPMQKENG 540 RGKARGVNPQ ALIWLTECSG IQLLGNSVVD PGPFLKVSVE ATATATGTGF TDGVSVPPAN 600 R |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
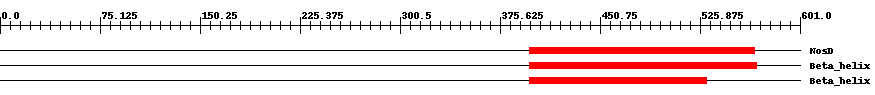
| pfam05048 | NosD | 0.0005 | 398 | 567 | 170 | + Periplasmic copper-binding protein (NosD). NosD is a periplasmic protein which is thought to insert copper into the exported reductase apoenzyme (NosZ). This region forms a parallel beta helix domain. | ||
| pfam13229 | Beta_helix | 3.0e-8 | 398 | 568 | 173 | + Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. | ||
| pfam13229 | Beta_helix | 2.0e-8 | 398 | 531 | 135 | + Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | ZP_01874996.1 | 6.99949e-42 | 37 | 531 | 10 | 509 | hypothetical protein LNTAR_21165 [Lentisphaera araneosa HTCC2155] |
| RefSeq | ZP_03723352.1 | 1.96182e-44 | 17 | 537 | 27 | 562 | hypothetical protein ObacDRAFT_9770 [Opitutaceae bacterium TAV2] |
| RefSeq | ZP_06241688.1 | 0 | 1 | 596 | 9 | 595 | hypothetical protein Vvad_PD3300 [Victivallis vadensis ATCC BAA-548] |
| RefSeq | ZP_06245028.1 | 0 | 33 | 594 | 21 | 581 | hypothetical protein Vvad_PD0287 [Victivallis vadensis ATCC BAA-548] |
| RefSeq | ZP_06245129.1 | 9.80909e-45 | 28 | 561 | 18 | 549 | Alpha-galactosidase [Victivallis vadensis ATCC BAA-548] |