

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_158718g0050 |
| Family | GT4 |
| Protein Properties | Length: 389 Molecular Weight: 43594.7 Isoelectric Point: 8.6425 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT4 | 211 | 337 | 4.8e-25 |
| EFFLVVGTHEPRKNIRTVIQAYSQLTSRERARCPLVLAGGRGWGKLDLPEPTEAMVRNGSVRFVHDMTDPQLRNLYEGARLLLAPSLYEGFGMPVVEGLA CGTPVAHSADTAMDEISGSIGKRVGAL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 389 Download |
| MTGVANYTFN IVRSVLELEP NLRFAGFRDF DWRRFDLAEA EKIAFSERKR DDAETSSASS 60 LTRVRRRLLD DVAGLPGSRK AAAVYREIRK GRFAWSVQRQ SLDLFHAFNF RPLTDPGVPI 120 LPVIYDLSTF RHPEFHPPDR VKWLEPLRQT IERAPLVQTI SDFSRREIAE LFAYPLAQIL 180 VAPPAAAPLF APLGEERTQP GLDLLGLTYG EFFLVVGTHE PRKNIRTVIQ AYSQLTSRER 240 ARCPLVLAGG RGWGKLDLPE PTEAMVRNGS VRFVHDMTDP QLRNLYEGAR LLLAPSLYEG 300 FGMPVVEGLA CGTPVAHSAD TAMDEISGSI GKRVGALDVD GWTDLLKASI NTTDHTDPTG 360 RAARIARAHE FDWRRSASLV IDAYRRIVS |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
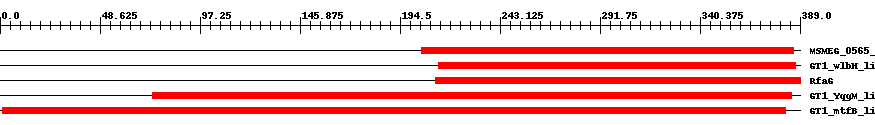
| TIGR04047 | MSMEG_0565_glyc | 7.0e-15 | 205 | 386 | 192 | + glycosyltransferase, MSMEG_0565 family. A conserved gene cluster found sporadically from Actinobacteria to Proteobacteria to Cyanobacteria features a radical SAM protein, an N-acetyltransferase, an oxidoreductase, and two additional proteins whose functional classes are unclear. The metabolic role of the cluster is probably biosynthetic. This glycosyltransferase, named from member MSMEG_0565 from Mycobacterium smegmatis, occurs in most but not all instances of the cluster [Unknown function, Enzymes of unknown specificity]. | ||
| cd03798 | GT1_wlbH_like | 1.0e-16 | 213 | 387 | 182 | + This family is most closely related to the GT1 family of glycosyltransferases. wlbH in Bordetella parapertussis has been shown to be required for the biosynthesis of a trisaccharide that, when attached to the B. pertussis lipopolysaccharide (LPS) core (band B), generates band A LPS. | ||
| COG0438 | RfaG | 1.0e-20 | 212 | 389 | 180 | + Glycosyltransferase [Cell envelope biogenesis, outer membrane] | ||
| cd03801 | GT1_YqgM_like | 5.0e-22 | 74 | 385 | 328 | + This family is most closely related to the GT1 family of glycosyltransferases and named after YqgM in Bacillus licheniformis about which little is known. Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. This group of glycosyltransferases is most closely related to the previously defined glycosyltransferase family 1 (GT1). The members of this family may transfer UDP, ADP, GDP, or CMP linked sugars. The diverse enzymatic activities among members of this family reflect a wide range of biological functions. The protein structure available for this family has the GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. The members of this family are found mainly in certain bacteria and archaea. | ||
| cd03809 | GT1_mtfB_like | 3.0e-63 | 1 | 382 | 389 | + This family is most closely related to the GT1 family of glycosyltransferases. mtfB (mannosyltransferase B) in E. coli has been shown to direct the growth of the O9-specific polysaccharide chain. It transfers two mannoses into the position 3 of the previously synthesized polysaccharide. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAA07751.1 | 0 | 1 | 385 | 14 | 379 | mannosyltransferase B [Escherichia coli] |
| GenBank | EFF13335.1 | 0 | 1 | 385 | 14 | 379 | conserved hypothetical protein [Escherichia coli B354] |
| RefSeq | YP_002387504.1 | 0 | 1 | 385 | 18 | 383 | mannosyltransferase B [Escherichia coli IAI1] |
| RefSeq | YP_578013.1 | 0 | 1 | 388 | 29 | 417 | glycosyl transferase, group 1 [Nitrobacter hamburgensis X14] |
| RefSeq | ZP_04629420.1 | 0 | 1 | 387 | 14 | 377 | Glycosyl transferase group 1 [Yersinia bercovieri ATCC 43970] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3l01_B | 0.000002 | 195 | 384 | 236 | 427 | A Chain A, Crystal Structure Of Monomeric Glycogen Synthase From Pyrococcus Abyssi |
| PDB | 3l01_A | 0.000002 | 195 | 384 | 236 | 427 | A Chain A, Crystal Structure Of Monomeric Glycogen Synthase From Pyrococcus Abyssi |
| PDB | 3fro_C | 0.000002 | 195 | 384 | 236 | 427 | A Chain A, Crystal Structure Of Pyrococcus Abyssi Glycogen Synthase With Open And Closed Conformations |
| PDB | 3fro_B | 0.000002 | 195 | 384 | 236 | 427 | A Chain A, Crystal Structure Of Pyrococcus Abyssi Glycogen Synthase With Open And Closed Conformations |
| PDB | 3fro_A | 0.000002 | 195 | 384 | 236 | 427 | A Chain A, Crystal Structure Of Pyrococcus Abyssi Glycogen Synthase With Open And Closed Conformations |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
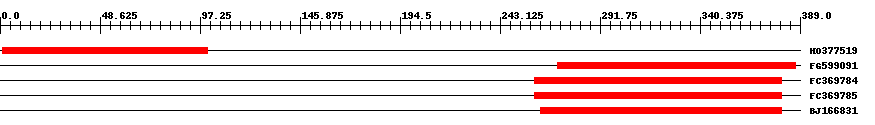 | ||||
| Hit | Length | Start | End | EValue |
| HO377519 | 101 | 1 | 101 | 2e-30 |
| FG599091 | 117 | 271 | 387 | 0.000000000001 |
| FC369784 | 125 | 260 | 380 | 0.0002 |
| FC369785 | 125 | 260 | 380 | 0.0002 |
| BJ166831 | 121 | 263 | 380 | 0.002 |
| Orthologous Group | |||||
|---|---|---|---|---|---|
| Species | ID | ||||
| Ricinus communis | 55201.m000024 | ||||
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |