

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_158983g0180 |
| Family | GH108 |
| Protein Properties | Length: 932 Molecular Weight: 96683.6 Isoelectric Point: 7.0324 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH108 | 735 | 818 | 1e-32 |
| TRLLKDEGGYTDHPSDPGGPTNFGITLADARRYWKGNATADDVRTMPQSAARKIYRERYWNAMRCDELPAGVDYAVFDYGVNSG | |||
| Full Sequence |
|---|
| Protein Sequence Length: 932 Download |
| MLNFKNLFRP PEAKASRTAQ VLAFESGGRA RWTPRDYAAL AREGYLANAI VHRAVRLIAE 60 NAASCGFLLY EGAQEREAHP LLQLLTRPNA RQDGASFFEA LYAHMLLAGN AYVEAVALGD 120 EVRELYTLRP DRMKVVPGHD GWAEAYEYSV AGRSVRFDQL ASSVPPILHL TFFHPLDDHY 180 GLAPIEAAAV AVDAVVQIGV LDADRTAPPT TPAEGDRHII ASGATGAWAG HADAVAAYQD 240 GAWRFLVPKP GWCAWCDADG ALLVYDGTAW TDVAGGAGAM PGSVAQLGVN DTASAPNLLT 300 VKSNAALFNA VSVAGGGTGH MRVQISKEAS AKTASVVFSD AFSGRAEFGL IGSDDFKLKV 360 SSDGSTFVEA LAIDQSSGNA ALPRGLSLTG VISPSQITAN QNDYNPSGLG AVSVLNLSSD 420 TLRSMSGLAG GAEGRVIALI NTGSQIISLL NESASSTAAN RFALGTDLTI AGKQAALLRY 480 DGTAARWRAM SRPGGRETLA ASRTYYVRTD GSDSNDGLSN ASGGAFLTIQ KAISAVAGLD 540 LNGKAVTIQV GNGTYAAGVS VTSPFVGGVP VLQGDTTTPG NVVISVSGDA VSVSNGAELG 600 IGGFKLVTAT AGSGLNATKA GRINVTGKME FGTCATAHMH SSYAGQIAVS ADYTISGGSL 660 YHWWSETAGG SIAVIGRTVT LTGTPAFTAF ANATIVAQIV AVSNTRDIAR FDRGNDSDRR 720 AVQLMAAESY NAALTRLLKD EGGYTDHPSD PGGPTNFGIT LADARRYWKG NATADDVRTM 780 PQSAARKIYR ERYWNAMRCD ELPAGVDYAV FDYGVNSGIG RAGKVLRRVL EMPDSTSAVT 840 NDVIAATSKR NAGDLVAAIC AERLAFLQGL KIFPVFGRGW TARVNGVRVA ALAMAQGRTP 900 ATATSAETSA GKAVVPVKNP AGPASAGAVI AA |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
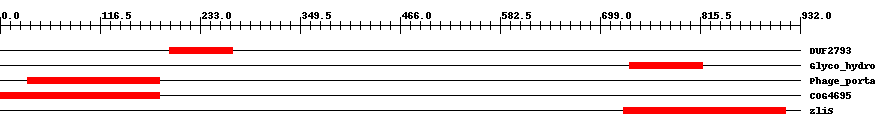 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam10983 | DUF2793 | 4.0e-20 | 197 | 271 | 75 | + Protein of unknown function (DUF2793). This is a bacterial family of proteins with unknown function. | ||
| pfam05838 | Glyco_hydro_108 | 2.0e-27 | 733 | 818 | 86 | + Glycosyl hydrolase 108. This family acts as a lysozyme (N-acetylmuramidase), EC:3.2.1.17. It contains a conserved EGGY motif near the N-terminus, the glutamic acid within this motif is essential for catalytic activity. In bacteria, it may activate the secretion of large proteins via the breaking and rearrangement of the peptidoglycan layer during secretion. It is frequently found at the N-terminus of proteins containing a C-terminal pfam09374 domain. | ||
| pfam04860 | Phage_portal | 2.0e-30 | 32 | 186 | 161 | + Phage portal protein. Bacteriophage portal proteins form a dodecamer and is located at a five-fold vertex of the viral capsid. The portal complex forms a channel through which the viral DNA is packaged into the capsid, and exits during infection. The portal protein is though to rotate during DNA packaging. Portal proteins from different phage show little sequence homology, so this family does not represent all portal proteins. | ||
| COG4695 | COG4695 | 3.0e-35 | 1 | 186 | 195 | + Phage-related protein [Function unknown] | ||
| COG3926 | zliS | 5.0e-40 | 726 | 915 | 194 | + Lysozyme family protein [General function prediction only] | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | YP_002289586.1 | 0 | 1 | 193 | 1 | 194 | phage portal protein, HK97 family [Oligotropha carboxidovorans OM5] |
| RefSeq | YP_317796.1 | 0 | 192 | 501 | 30 | 352 | hypothetical protein Nwi_1182 [Nitrobacter winogradskyi Nb-255] |
| RefSeq | YP_487064.1 | 0 | 192 | 491 | 30 | 325 | hypothetical protein RPB_3457 [Rhodopseudomonas palustris HaA2] |
| RefSeq | YP_780849.1 | 0 | 192 | 491 | 30 | 326 | hypothetical protein RPE_1925 [Rhodopseudomonas palustris BisA53] |
| RefSeq | ZP_06359547.1 | 0 | 192 | 487 | 30 | 323 | hypothetical protein Rpdx1DRAFT_3027 [Rhodopseudomonas palustris DX-1] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2is5_D | 4e-27 | 726 | 884 | 2 | 155 | A Chain A, Crystal Structure Of 3 Residues Truncated Version Of Protein Nmb1012 From Neisseria Meningitides |
| PDB | 2is5_C | 4e-27 | 726 | 884 | 2 | 155 | A Chain A, Crystal Structure Of 3 Residues Truncated Version Of Protein Nmb1012 From Neisseria Meningitides |
| PDB | 2is5_B | 4e-27 | 726 | 884 | 2 | 155 | A Chain A, Crystal Structure Of 3 Residues Truncated Version Of Protein Nmb1012 From Neisseria Meningitides |
| PDB | 2is5_A | 4e-27 | 726 | 884 | 2 | 155 | A Chain A, Crystal Structure Of 3 Residues Truncated Version Of Protein Nmb1012 From Neisseria Meningitides |
| PDB | 2ikb_D | 1e-26 | 727 | 884 | 2 | 154 | A Chain A, Crystal Structure Of A Protein Of Unknown Function Nmb1012 From Neisseria Meningitidis |