

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_159118g0670 |
| Family | PL12 |
| Protein Properties | Length: 1077 Molecular Weight: 116356 Isoelectric Point: 7.7291 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL12 | 392 | 531 | 8.7e-36 |
| GYTVSRGRMSGRESLAVFDHGPLGYLSIAAHGHADALSLTLHLDGRPVLVDPGTYLYHSGGSVRDRLRGTAVHNTLCINGTDQSEISGAFNWRRKATSEF TGLRETGHGVDVSAQHDGYARDFGLLCERTITVDASGLTV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1077 Download |
| MSIPELGHRL REAGKRRLSA TRSYDYSAFA IESADLPFLS VDIDRLQSID ADTRALWQSV 60 APTAKAPGIY RALGHDFHLS TEMDWHLDPV SGLRWPDDAY CFDIEYRHNR LRGDVKLVWE 120 LNRLQFVPVI AALSRLENDD ELAQSALDLI ESWIDANPPF KGINWISGIE LALRVVSILL 180 TIGLVGRERV GARLSRKIAA CLAAHAYWLK RFPSRYSSAN NHLIAEEAAL FVLGTLWQGA 240 EIDASAARAI LIAEAFRQIH DDGVGAEQSP TYTAFTCEWY MLAFTVAEAA GVPFPDNVIA 300 RVASAGEHLR WLLDSSGNHP RIGDDDEGRV IASGAGHEPL YIGSVVNGIA AVTGRQDLAA 360 DYGAPQLRNL LVGWPDATSV SGAGTRIFER GGYTVSRGRM SGRESLAVFD HGPLGYLSIA 420 AHGHADALSL TLHLDGRPVL VDPGTYLYHS GGSVRDRLRG TAVHNTLCIN GTDQSEISGA 480 FNWRRKATSE FTGLRETGHG VDVSAQHDGY ARDFGLLCER TITVDASGLT VRDKLLATGD 540 QASKPLTSIS IGFLLHPECS ARIEGATVVV TRGGEALMQV SAQPNVSVTL EGAEYSPAFG 600 VMESAARIVF PGSGAGLEFR RRRSPMKNLN VVLYFGANLV PRLSTFVLLL VLTRLLSMDD 660 YGMFALIVVT GEILDMATGG WVRLFVLSSE SKDRVPSAGR LGRALVLTVA SCVLSLLGAL 720 LLAIVQPSLS GWFTVAVAVY VLAFAVLRLG LTLLQTQQRH GLYAGVEIAR GILSLGCGVG 780 AVLAIEPTFF AASLGISLVT LVLGLLACAV AFRDLPWPYF PRQGYRAALL FGVPIVSVTL 840 LAQTVGWLDR FILNHALGPA SVGLYAAAYA LARQPVELFS GSLNPYLFPM LVRSYSSEGR 900 EAAGRVQSGN ILTLILLCGA VAVGIALLAQ PFAELVLPEA YRVEAARVMP WIAFATLFAS 960 LKNFCFDNAF YITKKNWQQF VSMAPPAVIS LLLGLLLIPQ GGPQRISLHT PVAIYCSVAR 1020 HCRASSFIWA GLSRDPRHPR ASGRAIATHH IVRGNRDIHR GLCINALCIR ICTGADA 1080 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| cd13128 | MATE_Wzx_like | 1.0e-14 | 768 | 966 | 205 | + Wzx, a subfamily of the multidrug and toxic compound extrusion (MATE)-like proteins. Escherichia coli Wzx and related proteins from other gram-negative bacteria are thought to act as flippases, assisting in the membrane translocation of lipopolysaccharides including those containing O-antigens. Proteins from the MATE family are involved in exporting metabolites across the cell membrane and are often responsible for multidrug resistance (MDR). | ||
| COG2244 | RfbX | 6.0e-17 | 763 | 1002 | 242 | + Membrane protein involved in the export of O-antigen and teichoic acid [General function prediction only] | ||
| pfam07940 | Hepar_II_III | 1.0e-31 | 391 | 535 | 145 | + Heparinase II/III-like protein. This family features sequences that are similar to a region of the Flavobacterium heparinum proteins heparinase II and heparinase III. The former is known to degrade heparin and heparan sulphate, whereas the latter predominantly degrades heparan sulphate. Both are secreted into the periplasmic space upon induction with heparin. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_772939.1 | 0 | 1 | 612 | 1 | 620 | hypothetical protein blr6299 [Bradyrhizobium japonicum USDA 110] |
| RefSeq | YP_001208842.1 | 0 | 79 | 600 | 92 | 625 | hypothetical protein BRADO7043 [Bradyrhizobium sp. ORS278] |
| RefSeq | YP_002782996.1 | 0 | 76 | 580 | 94 | 594 | hypothetical protein ROP_58040 [Rhodococcus opacus B4] |
| RefSeq | YP_592867.1 | 0 | 71 | 542 | 67 | 541 | heparinase II/III-like [Candidatus Koribacter versatilis Ellin345] |
| RefSeq | ZP_04242214.1 | 0 | 81 | 510 | 79 | 499 | hypothetical protein bcere0018_49180 [Bacillus cereus Rock1-15] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 4fnv_A | 0.00000005 | 59 | 499 | 100 | 544 | A Chain A, Crystal Structure Of Heparinase Iii |
| Transmembrane Domains | |||||
|---|---|---|---|---|---|
 | |||||
| Start | End | ||||
| 164 | 186 | ||||
| 631 | 653 | ||||
| 663 | 685 | ||||
| 705 | 724 | ||||
| 729 | 751 | ||||
| 763 | 785 | ||||
| 790 | 812 | ||||
| 825 | 847 | ||||
| 851 | 873 | ||||
| 911 | 933 | ||||
| 948 | 966 | ||||
| 979 | 998 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
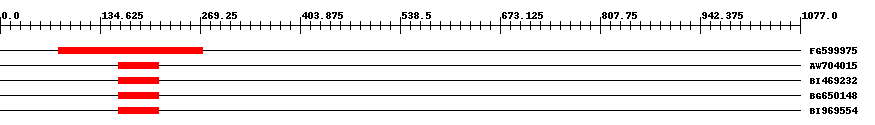 | ||||
| Hit | Length | Start | End | EValue |
| FG599975 | 200 | 79 | 272 | 0.00000000005 |
| AW704015 | 59 | 159 | 214 | 0.52 |
| BI469232 | 60 | 159 | 214 | 1.1 |
| BG650148 | 60 | 159 | 214 | 1.2 |
| BI969554 | 59 | 159 | 214 | 1.2 |
| Orthologous Group | |||||
|---|---|---|---|---|---|
| Species | ID | ||||
| Physcomitrella patens | Pp1s2660_1V6.1 | ||||
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |