

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_208475g0010 |
| Family | AA1 |
| Protein Properties | Length: 706 Molecular Weight: 77856.9 Isoelectric Point: 8.0417 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA1 | 20 | 234 | 0 |
| CPTIHARNGDTVIVKVYNNAQYNATIHWHGVRQFRTGWSDGPEFITQCPIRPGGSYTYKFTLTDQEGTLWWHGHSSWLRATVYGALIISPRLGSTYPYTK PHGQVPILLGEWWNRNPIDVVNQATRTGAAPSVSDAFTINGRPGDLYQCSSSDTLHISVKNGETKLLRVINAALNTDLFFSVGSHTMTVVAVDALYTKPF QTNVLMLGPGQTTDL | |||
| AA1 | 235 | 509 | 0 |
| QSTTVKRLCGTRNIITVNGQFPGPTIYARNGDTVIVKVYNNAQYNATIHWHGVRQFRTGWSDGPEFITQCPICPGGSYTYKFTLTDQEGTLWWHGHSSWL RATVYGALIISPRLGTTYPYYKPHGQLPILLGEWWNRNPIDVVNQATRTGAAPNVSDAFTINGRPGDLYQCSSSDTLRVSVKNGETKLLRVINAALNTDL FFSVGCHTMTVVAVDALYTKPFQTNVLMLGPGQTTDVLVTADKATGRYYMAARAYSSGQGVPFDNTTTVAILEYE | |||
| Full Sequence |
|---|
| Protein Sequence Length: 706 Download |
| MPHVVDCYGL REATYSDIRC PTIHARNGDT VIVKVYNNAQ YNATIHWHGV RQFRTGWSDG 60 PEFITQCPIR PGGSYTYKFT LTDQEGTLWW HGHSSWLRAT VYGALIISPR LGSTYPYTKP 120 HGQVPILLGE WWNRNPIDVV NQATRTGAAP SVSDAFTING RPGDLYQCSS SDTLHISVKN 180 GETKLLRVIN AALNTDLFFS VGSHTMTVVA VDALYTKPFQ TNVLMLGPGQ TTDLQSTTVK 240 RLCGTRNIIT VNGQFPGPTI YARNGDTVIV KVYNNAQYNA TIHWHGVRQF RTGWSDGPEF 300 ITQCPICPGG SYTYKFTLTD QEGTLWWHGH SSWLRATVYG ALIISPRLGT TYPYYKPHGQ 360 LPILLGEWWN RNPIDVVNQA TRTGAAPNVS DAFTINGRPG DLYQCSSSDT LRVSVKNGET 420 KLLRVINAAL NTDLFFSVGC HTMTVVAVDA LYTKPFQTNV LMLGPGQTTD VLVTADKATG 480 RYYMAARAYS SGQGVPFDNT TTVAILEYEG RQSCGGPNGS RFAASMNNIS FVLPTTSSIL 540 QAQHFGVKGV FSGDFPDNPP VQFDYTAQNV NRDLWSPVKG TKVKVLNYNT TVQVILQGTN 600 IFAGESHPIH LHGYDFYIVG TGFGNYNPQN DPQKFNLVDP PMRNTVNVPV NGWAAIRFVA 660 DNPGAWLLHC HLDAHITWGL AMVFLVNNGP NSLLSLESPP VDLPQC |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR03388 | ascorbase | 6.0e-31 | 21 | 231 | 234 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| PLN02604 | PLN02604 | 8.0e-79 | 243 | 684 | 527 | + oxidoreductase | ||
| TIGR03388 | ascorbase | 7.0e-83 | 243 | 684 | 517 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| TIGR03389 | laccase | 2.0e-119 | 21 | 241 | 221 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| TIGR03389 | laccase | 0 | 231 | 706 | 535 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK37824.1 | 0 | 21 | 234 | 66 | 281 | AF132120_1 laccase [Pinus taeda] |
| GenBank | AAK37824.1 | 0 | 234 | 706 | 42 | 576 | AF132120_1 laccase [Pinus taeda] |
| GenBank | ABR17528.1 | 0 | 21 | 234 | 62 | 276 | unknown [Picea sitchensis] |
| GenBank | ABR17528.1 | 0 | 234 | 706 | 38 | 570 | unknown [Picea sitchensis] |
| RefSeq | NP_001044679.1 | 0 | 234 | 706 | 33 | 567 | Os01g0827300 [Oryza sativa (japonica cultivar-group)] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1asq_B | 0 | 243 | 684 | 19 | 521 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1asq_B | 1e-24 | 21 | 238 | 34 | 268 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1asq_A | 0 | 243 | 684 | 19 | 521 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1asq_A | 1e-24 | 21 | 238 | 34 | 268 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1asp_B | 0 | 243 | 684 | 19 | 521 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DV974494 | 308 | 270 | 575 | 0 |
| DV974494 | 204 | 33 | 234 | 0 |
| CO473376 | 218 | 230 | 447 | 0 |
| CO473376 | 190 | 21 | 210 | 0 |
| CO474791 | 206 | 246 | 451 | 0 |
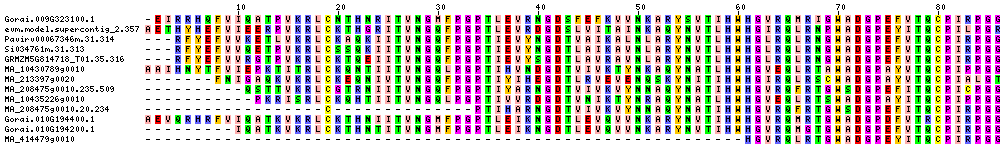
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |