

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000118804 |
| Family | GT5 |
| Protein Properties | Length: 880 Molecular Weight: 99418.7 Isoelectric Point: 5.0075 |
| Chromosome | Chromosome/Scaffold: 000149326 Start: 7958 End: 13713 |
| Description | starch synthase 4 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 567 | 726 | 2.8026e-45 |
| MAPLYWDLYAPKGLNSGRICFTCHNFEYQGTARASELASCGLDVHQLNRPDRMQDNSAHDRINAVKGAVVFSNIVTTVSPTYAQEVRTAEGGHGLHSTLN FHSKKFVGILNGIDADAWNPATDAYLKVQYXANDRQGKAENKEALRRILRLSSADVKRPL | |||
| GT5 | 728 | 869 | 0 |
| EFEGIASHFANHDHIRLILKYDDSLSHTIYAASDMFIIPSIFEPCGLTQMIAMRYGSIPIARKTGGLNDSVFDVDDDTVPLQFRNGYSFLTPDEQGLNGA MERAFDLYXNNPDXWQQLVQKVMNIDFSWDTSASQYEELYSK | |||
| Full Sequence |
|---|
| Protein Sequence Length: 880 Download |
| MAVRLTTWFV TQGLTISGSN STHCSNSSSR FSFHCTPASS CKLRHRXLSC YCVNKRQRLK 60 KKDSVEQSST TADFQLNGDD DSESENASSA GVVVPIESVS DDEARTANGD DSISTALTPS 120 DEANPSAYST QDLVDMIRNA EKNIHILNQA RVNALEDLDK ILGEKEALQG EMNALEMRLA 180 ETDARIRVAA QEKIKVELLE NQLDEMRNDL NLNSGGGVER GQVEIFENEX ELFNEEAPVP 240 YRTSINALVT NLNALRLENQ SLRSDVEALR EELSYVKNTD ERVVMLEKQR STLESALKEL 300 ELKLSVSQED VSKLSNLKVE CKGLWEKVES LQLLLDKSTK QADQAITVLQ QNQEIRKKVD 360 KLEESLETAN IYKESSEKMQ QYNELMQQKI KLMEDRLQRS DEEIHSYVQL YQESVEEFQD 420 TLNTLKEESK RRAVDEPVDD MPWEFWSRLL LMIDGWLFEK KISMDDAKVL REMVWKRDRR 480 LRDSYMACKE KNVNEAVSTF LKLISSQTSP GLHVVHIAAE MAPVAKVGGL GDVVAGLGKA 540 LQKKGHLVEI ILPKYDCMQY DRVPDLMAPL YWDLYAPKGL NSGRICFTCH NFEYQGTARA 600 SELASCGLDV HQLNRPDRMQ DNSAHDRINA VKGAVVFSNI VTTVSPTYAQ EVRTAEGGHG 660 LHSTLNFHSK KFVGILNGID ADAWNPATDA YLKVQYXAND RQGKAENKEA LRRILRLSSA 720 DVKRPLREFE GIASHFANHD HIRLILKYDD SLSHTIYAAS DMFIIPSIFE PCGLTQMIAM 780 RYGSIPIARK TGGLNDSVFD VDDDTVPLQF RNGYSFLTPD EQGLNGAMER AFDLYXNNPD 840 XWQQLVQKVM NIDFSWDTSA SQYEELYSKS VARARVAART |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR02095 | glgA | 1.0e-5 | 512 | 564 | 53 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 5.0e-113 | 567 | 869 | 343 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| TIGR02095 | glgA | 7.0e-117 | 567 | 870 | 343 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| PLN02939 | PLN02939 | 0 | 567 | 870 | 346 | + transferase, transferring glycosyl groups | ||
| PLN02939 | PLN02939 | 0 | 38 | 568 | 543 | + transferase, transferring glycosyl groups | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | EEC71396.1 | 0 | 240 | 880 | 113 | 830 | hypothetical protein OsI_03547 [Oryza sativa Indica Group] |
| GenBank | EEE55309.1 | 0 | 28 | 880 | 12 | 930 | hypothetical protein OsJ_03283 [Oryza sativa Japonica Group] |
| RefSeq | XP_002519725.1 | 2e-21 | 1 | 188 | 1 | 229 | starch synthase, putative [Ricinus communis] |
| RefSeq | XP_002519725.1 | 0 | 324 | 568 | 312 | 556 | starch synthase, putative [Ricinus communis] |
| RefSeq | XP_002519725.1 | 0 | 566 | 877 | 641 | 994 | starch synthase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1rzv_B | 7.00649e-44 | 587 | 874 | 160 | 479 | A Chain A, Crystal Structure Of The Glycogen Synthase From Agrobacterium Tumefaciens (Non-Complexed Form) |
| PDB | 1rzv_A | 7.00649e-44 | 587 | 874 | 160 | 479 | A Chain A, Crystal Structure Of The Glycogen Synthase From Agrobacterium Tumefaciens (Non-Complexed Form) |
| PDB | 1rzu_B | 7.00649e-44 | 587 | 874 | 160 | 479 | A Chain A, Crystal Structure Of The Glycogen Synthase From A. Tumefaciens In Complex With Adp |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
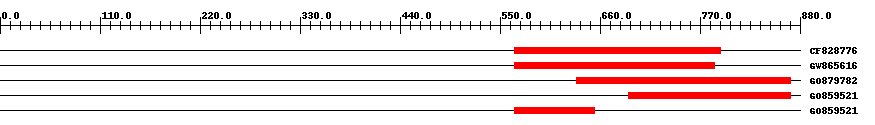 | ||||
| Hit | Length | Start | End | EValue |
| CF828776 | 270 | 566 | 793 | 0 |
| GW865616 | 263 | 566 | 786 | 0 |
| GO879782 | 278 | 634 | 869 | 0 |
| GO859521 | 222 | 691 | 870 | 0 |
| GO859521 | 89 | 566 | 654 | 0 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
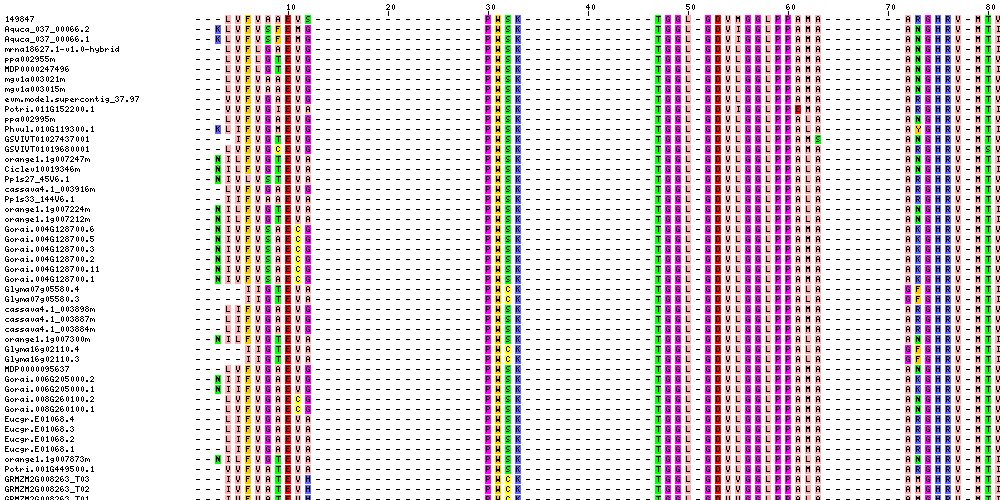 |