

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000212739 |
| Family | CBM22 |
| Protein Properties | Length: 1108 Molecular Weight: 122254 Isoelectric Point: 6.7462 |
| Chromosome | Chromosome/Scaffold: 016467170 Start: 5166 End: 10509 |
| Description | glycosyl hydrolase family 10 protein / carbohydrate-binding domain-containing protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM22 | 225 | 359 | 2.4e-30 |
| NIIMNHDFSGGLHSWHPNCCNGFVASADSGHPEVKAGNYAVVTNRKESWQGLEQDITRRISPGSTYSVSACVGVCGSLQGSADVLATLKLEYRGSATNHL QVGRCSVSKGRWGNLDGKFSLSTMPDRVVFYLEGP | |||
| CBM22 | 64 | 190 | 1.6e-31 |
| NIVLNHDFSGGLHSWHXNHCNGFVVDSAGSYAVVTNRQQCWQGLEQDITGRISPGNTYSVSARVGVSGTLQGSADVLATLKLESRGSATSYMRIGRSSVS NGKWESLDGKFSLSTMPDRVVFYLEGP | |||
| CBM22 | 394 | 530 | 9.3e-33 |
| NIILNPNFEDALNNWSGRGCKIVLHDSMGDGKIVPQSGKVFAAATERTQSWNGIQQDITGRVQRKLAYEATAVVRIFGNNVTXAVVRATLWVQSPNQREQ YIGIANVQATDKDWTQLRGKFLLNGSPSKVVVYLEGP | |||
| CBM22 | 565 | 706 | 2.79979e-42 |
| NIIENSNLSNXTNGWFPLGNCTLSVTTGSPHILPPMARESLGPHEPLSGRYILVTKRTQTWMGPAQMIGDKLKLFLTYQVSAWVRIGAGATGPQNVNVAL SVDNQWVNGXQAEVSDTRWHEIGGSFRVEKQPSKVMVYIQGP | |||
| GH10 | 763 | 1031 | 0 |
| KQTQNSFPIGTCISRTNIDNEDYVDFFIKNFNWAVFGNELKWYWTEPQKGNFNYKDADEMVDXCKSHNIEMRGHCIFWEVIDTVQQWIRSLSQSDLSTAV QNRLTDLLTRYKGKFRHYDVNNEMLHGSFYQDKLGKDIRANMFKNANQLDPSATLFVNDYHVEDGCDTRSSPEKYTDQILDLQQQGAPVGGIGIQGHIDS PVGPIVCTALDKLGILGLPIWFTELDVSSSNEYVRADDLEVMLREAFANPTVEGVMLWGFWELFMSREN | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1108 Download |
| MRRLIAWCFR NRVSDSSNHX KQKHPEGSXE AMDKKNQQTN NNAADNVEFL RQNVADSSSS 60 RGPNIVLNHD FSGGLHSWHX NHCNGFVVDS AGSYAVVTNR QQCWQGLEQD ITGRISPGNT 120 YSVSARVGVS GTLQGSADVL ATLKLESRGS ATSYMRIGRS SVSNGKWESL DGKFSLSTMP 180 DRVVFYLEGP PAGVDLHIKS VVISCSEGQS ENQNLANSSS SNATNIIMNH DFSGGLHSWH 240 PNCCNGFVAS ADSGHPEVKA GNYAVVTNRK ESWQGLEQDI TRRISPGSTY SVSACVGVCG 300 SLQGSADVLA TLKLEYRGSA TNHLQVGRCS VSKGRWGNLD GKFSLSTMPD RVVFYLEGPS 360 PGVDLLIKSV LICSLSPNEW QSGSTGNFND GEENIILNPN FEDALNNWSG RGCKIVLHDS 420 MGDGKIVPQS GKVFAAATER TQSWNGIQQD ITGRVQRKLA YEATAVVRIF GNNVTXAVVR 480 ATLWVQSPNQ REQYIGIANV QATDKDWTQL RGKFLLNGSP SKVVVYLEGP LAGTDILVNS 540 FVVKHAEKVP PSPPPVIEFS AFGVNIIENS NLSNXTNGWF PLGNCTLSVT TGSPHILPPM 600 ARESLGPHEP LSGRYILVTK RTQTWMGPAQ MIGDKLKLFL TYQVSAWVRI GAGATGPQNV 660 NVALSVDNQW VNGXQAEVSD TRWHEIGGSF RVEKQPSKVM VYIQGPAAGV DLMVAGLQIF 720 PVDRPARFRH LKRQTDKVRK CDIVLKFSGL DSSSMLGTFV KVKQTQNSFP IGTCISRTNI 780 DNEDYVDFFI KNFNWAVFGN ELKWYWTEPQ KGNFNYKDAD EMVDXCKSHN IEMRGHCIFW 840 EVIDTVQQWI RSLSQSDLST AVQNRLTDLL TRYKGKFRHY DVNNEMLHGS FYQDKLGKDI 900 RANMFKNANQ LDPSATLFVN DYHVEDGCDT RSSPEKYTDQ ILDLQQQGAP VGGIGIQGHI 960 DSPVGPIVCT ALDKLGILGL PIWFTELDVS SSNEYVRADD LEVMLREAFA NPTVEGVMLW 1020 GFWELFMSRE NSHLVNAEGD INEAGKRFLE LKQEWLSHAH GHIDQQGEFG FRGFPGTYSV 1080 EVFTASKKPA KTFVVDKGES PVEVSIAL |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
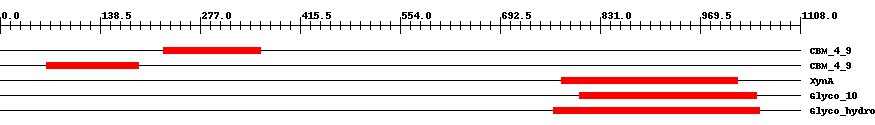 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam02018 | CBM_4_9 | 2.0e-15 | 226 | 361 | 138 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| pfam02018 | CBM_4_9 | 1.0e-17 | 64 | 192 | 135 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| COG3693 | XynA | 1.0e-32 | 777 | 1022 | 267 | + Beta-1,4-xylanase [Carbohydrate transport and metabolism] | ||
| smart00633 | Glyco_10 | 2.0e-76 | 802 | 1048 | 268 | + Glycosyl hydrolase family 10. | ||
| pfam00331 | Glyco_hydro_10 | 1.0e-78 | 767 | 1052 | 306 | + Glycosyl hydrolase family 10. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| GO:0016798 | hydrolase activity, acting on glycosyl bonds |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAX33301.1 | 0 | 38 | 538 | 4 | 521 | putative endo-1,4-beta-xylanase [Populus tremula x Populus tremuloides] |
| GenBank | AAX33301.1 | 0 | 209 | 1108 | 14 | 915 | putative endo-1,4-beta-xylanase [Populus tremula x Populus tremuloides] |
| RefSeq | XP_002301133.1 | 0 | 50 | 538 | 150 | 655 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 64 | 1108 | 2 | 1049 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 393 | 719 | 1 | 314 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1nq6_A | 5e-29 | 771 | 1043 | 15 | 291 | A Chain A, Crystal Structure Of The Catalytic Domain Of Xylanase A From Streptomyces Halstedii Jm8 |
| PDB | 1i1x_A | 2e-28 | 792 | 1021 | 39 | 268 | A Chain A, Crystal Structure Of The Catalytic Domain Of Xylanase A From Streptomyces Halstedii Jm8 |
| PDB | 1i1w_A | 2e-28 | 792 | 1021 | 39 | 268 | A Chain A, 0.89a Ultra High Resolution Structure Of A Thermostable Xylanase From Thermoascus Aurantiacus |
| PDB | 2bnj_A | 2e-27 | 792 | 1021 | 39 | 268 | A Chain A, The Xylanase Ta From Thermoascus Aurantiacus Utilizes Arabinose Decorations Of Xylan As Significant Substrate Specificity Determinants |
| PDB | 1xyz_B | 3e-27 | 771 | 1022 | 39 | 307 | A Chain A, A Common Protein Fold And Similar Active Site In Two Distinct Families Of Beta-Glycanases |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EB442796 | 298 | 592 | 889 | 0 |
| EX570690 | 294 | 815 | 1108 | 0 |
| EB442796 | 125 | 248 | 372 | 0.00000002 |
| EB442796 | 113 | 91 | 203 | 0.00000007 |
| EB442796 | 120 | 419 | 538 | 0.0000004 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |