

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000247130 |
| Family | GH17 |
| Protein Properties | Length: 452 Molecular Weight: 49324.7 Isoelectric Point: 8.4221 |
| Chromosome | Chromosome/Scaffold: 0007271180 Start: 1085 End: 6857 |
| Description | Glycosyl hydrolase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 167 | 242 | 8.2e-26 |
| VGVCYGRNGNNLPAEGEVVDLYKSNGIGRMRIYEPNEATLQALRGSNIELTVTILNXELXALNDAAAATAWVQKNV | |||
| GH17 | 244 | 451 | 0 |
| SLLPAIQNIHSAIVAANLQGQIKVSTAIDTTLVANPFPPSDGVYDAANQFIXPVIDFLVSSGAPLLVNVYPYFSYXDNPSIDLAYALFTSQGVVVPDGTR YPSLFDALLDAQYAALEKAGAPNMEIVVSESGWPSEGGDQATPQNAATFYQNLINHVTSTTGTPKRPGKAIETYLFAMFDENLKDGKPVEKHFGVFSPNK QPKYQLTF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 452 Download |
| MSWDLGIFRL KVIRVPSLRA KLEVLPLWLC LLQTPSRVPD PAVLRSRSVH LPISFKLNPR 60 PPMGPKLCYV RSFELGLYST RRGRICRMGG WGGRESXRVA GLWRWSNGFL PSRXFGWLVX 120 WAVXVMVALE ISGDGGDCTV VVVILSPRGN IDGIMGRFLI LNGAQSVGVC YGRNGNNLPA 180 EGEVVDLYKS NGIGRMRIYE PNEATLQALR GSNIELTVTI LNXELXALND AAAATAWVQK 240 NVGSLLPAIQ NIHSAIVAAN LQGQIKVSTA IDTTLVANPF PPSDGVYDAA NQFIXPVIDF 300 LVSSGAPLLV NVYPYFSYXD NPSIDLAYAL FTSQGVVVPD GTRYPSLFDA LLDAQYAALE 360 KAGAPNMEIV VSESGWPSEG GDQATPQNAA TFYQNLINHV TSTTGTPKRP GKAIETYLFA 420 MFDENLKDGK PVEKHFGVFS PNKQPKYQLT FG |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 5.0e-7 | 366 | 443 | 84 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 3.0e-128 | 167 | 451 | 312 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAX81589.1 | 0 | 159 | 452 | 1 | 320 | beta-1,3-glucanase [Fragaria x ananassa] |
| GenBank | AAX81590.1 | 0 | 165 | 452 | 33 | 346 | beta-1,3-glucanase [Fragaria x ananassa] |
| DDBJ | BAC66141.1 | 0 | 165 | 452 | 33 | 346 | beta-1,3-glucanase [Fragaria x ananassa] |
| DDBJ | BAC66185.1 | 0 | 165 | 452 | 33 | 346 | beta-1,3-glucanase [Fragaria x ananassa] |
| DDBJ | BAC66186.1 | 0 | 159 | 452 | 27 | 346 | beta-1,3-glucanase [Fragaria x ananassa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3f55_D | 0 | 167 | 451 | 2 | 315 | A Chain A, The Crystal Structure Of Rice (Oryza Sativa L.) Os4bglu12 |
| PDB | 3f55_C | 0 | 167 | 451 | 2 | 315 | A Chain A, The Crystal Structure Of Rice (Oryza Sativa L.) Os4bglu12 |
| PDB | 3f55_B | 0 | 167 | 451 | 2 | 315 | A Chain A, The Crystal Structure Of Rice (Oryza Sativa L.) Os4bglu12 |
| PDB | 3f55_A | 0 | 167 | 451 | 2 | 315 | A Chain A, The Crystal Structure Of Rice (Oryza Sativa L.) Os4bglu12 |
| PDB | 3em5_D | 0 | 167 | 451 | 2 | 315 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR992002 | 199 | 256 | 452 | 0 |
| CV657008 | 198 | 256 | 451 | 0 |
| HE645802 | 210 | 245 | 452 | 0 |
| FE969327 | 204 | 245 | 446 | 0 |
| HO779722 | 320 | 159 | 451 | 0 |
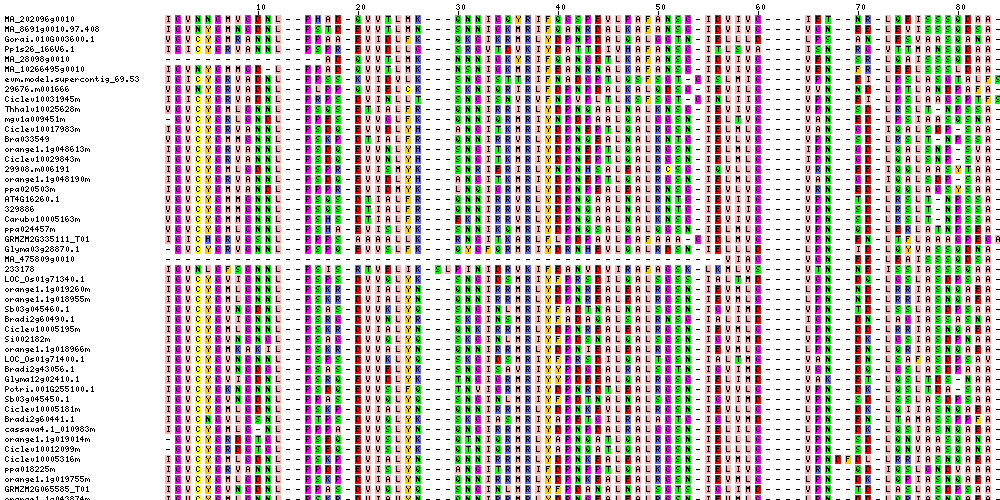
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |