

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000282861 |
| Family | GH17 |
| Protein Properties | Length: 426 Molecular Weight: 47450.6 Isoelectric Point: 4.5171 |
| Chromosome | Chromosome/Scaffold: 005813234 Start: 4294 End: 7695 |
| Description | beta-1,3-glucanase 5 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 1 | 91 | 1.9e-28 |
| MERVGGAGVEVAVTESGWPSAGNGIFTSPELAGTYNRNFMKHVTSLKGTPKRPGHYIEGFIFALFNENQKPAGVEQNFGLFYPTMKPVYTV | |||
| GH17 | 136 | 425 | 0 |
| IGVNYGTLGDNLPSHEDVVQLFKDNHINKMRLFNPNPEVLKALMGQGISIVLGIPNEDLASLSESQEAVDQWFAVNVQPYIDDIEFNYIAVGNEVIPGEL GSYVWQVMQYLQHTIDEYHYQSLKFSGEAAGIMASICPFLNYQGSPLMINVYPYFAYASKASDIPLDYAQFTATKPWLQDGNLSYYNLFDATVDAFFSAI EKSGGPDVAVVVSESGWPSAGNGDFTDTELAGTYVRNFVRHITSNVGTPKRPGAYIEGFVFAMFNENQKPSEEEQNFGIFYPNKKPVYPV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 426 Download |
| MERVGGAGVE VAVTESGWPS AGNGIFTSPE LAGTYNRNFM KHVTSLKGTP KRPGHYIEGF 60 IFALFNENQK PAGVEQNFGL FYPTMKPVYT VLYYKALSSD QKVMAVSQWI AFILSFLATI 120 YNHNIEIVEA KMTFDIGVNY GTLGDNLPSH EDVVQLFKDN HINKMRLFNP NPEVLKALMG 180 QGISIVLGIP NEDLASLSES QEAVDQWFAV NVQPYIDDIE FNYIAVGNEV IPGELGSYVW 240 QVMQYLQHTI DEYHYQSLKF SGEAAGIMAS ICPFLNYQGS PLMINVYPYF AYASKASDIP 300 LDYAQFTATK PWLQDGNLSY YNLFDATVDA FFSAIEKSGG PDVAVVVSES GWPSAGNGDF 360 TDTELAGTYV RNFVRHITSN VGTPKRPGAY IEGFVFAMFN ENQKPSEEEQ NFGIFYPNKK 420 PVYPVF |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 1.0e-5 | 148 | 419 | 300 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 7.0e-21 | 18 | 91 | 75 | + Glycosyl hydrolases family 17. | ||
| pfam00332 | Glyco_hydro_17 | 9.0e-97 | 136 | 425 | 312 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAN74958.1 | 4e-25 | 1 | 91 | 211 | 301 | hypothetical protein [Vitis vinifera] |
| EMBL | CAN74958.1 | 0 | 125 | 426 | 11 | 302 | hypothetical protein [Vitis vinifera] |
| RefSeq | NP_197534.1 | 8e-30 | 1 | 91 | 262 | 353 | BG5 (beta-1,3-glucanase 5); glucan 1,3-beta-glucosidase/ hydrolase, hydrolyzing O-glycosyl compounds [Arabidopsis thaliana] |
| RefSeq | NP_197534.1 | 0 | 112 | 426 | 11 | 354 | BG5 (beta-1,3-glucanase 5); glucan 1,3-beta-glucosidase/ hydrolase, hydrolyzing O-glycosyl compounds [Arabidopsis thaliana] |
| RefSeq | XP_002513812.1 | 0 | 136 | 425 | 22 | 333 | Lichenase precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 136 | 425 | 1 | 310 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 2cyg_A | 2e-23 | 1 | 93 | 222 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1ghr_A | 0 | 136 | 425 | 1 | 304 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1ghr_A | 2e-20 | 1 | 91 | 218 | 304 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1aq0_B | 0 | 136 | 425 | 1 | 304 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
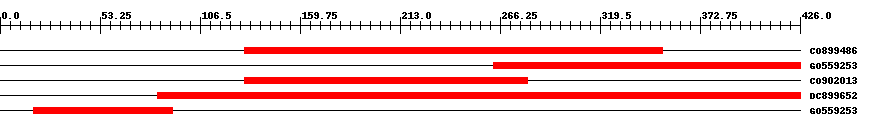 | ||||
| Hit | Length | Start | End | EValue |
| CO899486 | 244 | 130 | 353 | 0 |
| GO559253 | 164 | 263 | 426 | 0 |
| CO902013 | 172 | 130 | 281 | 0 |
| DC899652 | 376 | 84 | 426 | 0 |
| GO559253 | 75 | 18 | 92 | 3e-29 |
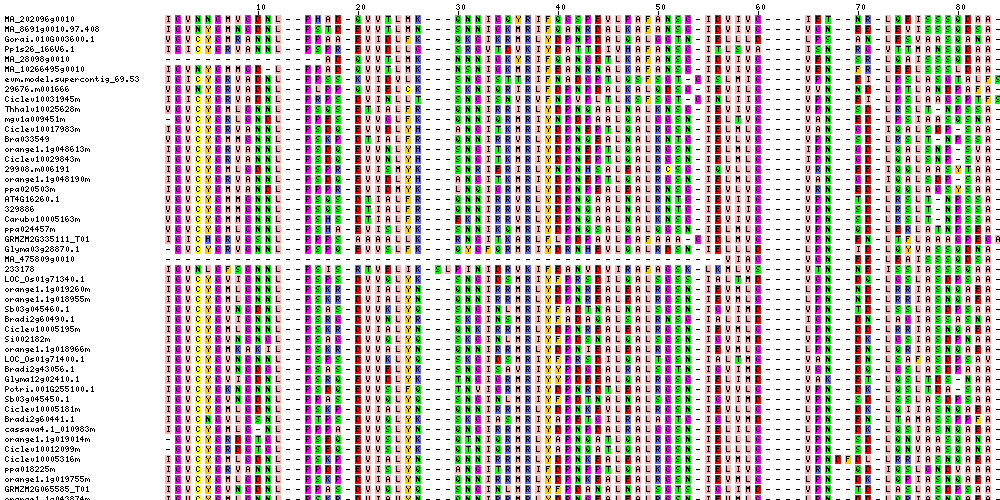
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |