

| Basic Information | |
|---|---|
| Species | Phaseolus vulgaris |
| Cazyme ID | Phvul.003G062900.1 |
| Family | CBM45 |
| Protein Properties | Length: 1457 Molecular Weight: 163484 Isoelectric Point: 6.7324 |
| Chromosome | Chromosome/Scaffold: 03 Start: 8679229 End: 8697331 |
| Description | Pyruvate phosphate dikinase, PEP/pyruvate binding domain |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM45 | 449 | 528 | 2.9e-21 |
| LHWALSRTSEEWLLPPGNSLPPGSVTMNEAAETPFKAGSLSHPSFEVQSLDIEVDDDTFKGIPFVILSEGKWIKNNGSNF | |||
| CBM45 | 120 | 195 | 3e-23 |
| LHWGVVCDQPGKWVLPSRRPEGTKVYKNKALRTPFMKADSESFLRIEIHDPAAQSIEFLILDEAKNKWFKNNGENF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1457 Download |
| MSQSIFHQTV LCQTQTVAEH QSKVSSFAVS VNKGKKNLGL RTSFRGNRLC VRKCKLAMGK 60 HRHVDAIPRA VLTTNPASEL SGRFILGGNI ELQVSVSSAQ PGAATQVDIK VSYSSGSLLL 120 HWGVVCDQPG KWVLPSRRPE GTKVYKNKAL RTPFMKADSE SFLRIEIHDP AAQSIEFLIL 180 DEAKNKWFKN NGENFHIKLP VKNKLSQEVS VPEDLVQIQA YLRWERKGKQ MYTPEQEKVE 240 YEAARQELLE EVSRGTSVQD LRARLTKNTK AAEVKEPSVS ETKTIPDELV QIQSYIRWEK 300 AGKPNYSQEQ QLMEFEEARK ELSAELEKGA SLDEIRKKII KGEVQTKVAK QLKTKTYFRA 360 ERIQRKNRDL RQIINRIVDE NIVEQFIDVP KSLTVIEHYA KEREENESGP VLNKTIYKLD 420 DNDLLVLVTK DAGKIKVHLA TNSKKPLTLH WALSRTSEEW LLPPGNSLPP GSVTMNEAAE 480 TPFKAGSLSH PSFEVQSLDI EVDDDTFKGI PFVILSEGKW IKNNGSNFYI EFAGKKQIRK 540 DFGDSKGTAK FLLDKIAEQE SEAQKSFMHR FNIASNLIDE AKSAGRLGLA GILVWMRFMA 600 TRQLIWNKNY NVKPREISKA QDRLTDLLQD VYASYPQYRE IVRMILSTVG RGGEGDVGQR 660 IRDEILVIQR NNDCKGGMME EWHQKLHNNT SPDDVVICQA LIDYIKNDFD TGVYWKTLND 720 NGITKERLLS YDRAIHSEPN FRRDQKEGLL RDLGNYMRTL KAVHSGADLE SAISNCMGYK 780 SEGQGFMVGV QINPVPGLPA GFQGLLEFVM EHVEDKNVEP LLEGLLEARE ELHPSLGKSQ 840 SRLKDLLFLD VALDSTVRTA VERGYEELNN AAPEKIMYFI CLVLENLSLS SDDNEDLIYC 900 LKGWDLALTK CKSNDTHWAL YAKSVLDRTR LALTNKAQLY QEILQPSAEY LGSLLGVDQW 960 AVEIFTEEII RAGSAASLST LLNRLDPVLR KTANLGSWQV ISPVETVGYV EVVDELLSVQ 1020 NKSYERPTIL IAKSVKGEEE IPDGTVAVLT PDMPDVLSHV SVRARNSKVC FATCFDPNIL 1080 ANLQESRGKL LRLKPTSADV VYSQVEEGEF IDDKSSHLKD VGSVSPISLV RKKFSGRYAV 1140 SSEEFTGEMV GAKSRNITYL KGKVASWIGI PTSVAIPFGV FEHVLSDKSN QAVAERVNIL 1200 KKKLIEGDFS VLKEIRETVL QLNAPPQLVE ELKSKMKSSG MPWPGDEGEQ RWEQAWKAIK 1260 KVWGSKWNER AYFSTRKVKL DHEYLSMAVL VQEVVNADYA FVIHTTNPSS GDSSEIYAEV 1320 VKGLGETLVG AYPGRALSFI CKKSDLNSPQ VLGYPSKPVG LFIRQSIIFR SDSNGEDLEG 1380 YAGAGLYDSV PMDEEEKVVL DYSSDQLMLD GSFRRTILSS IARAGNEIEG LYGSPQDIEG 1440 VIKDGKLYVV QTRPQM* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
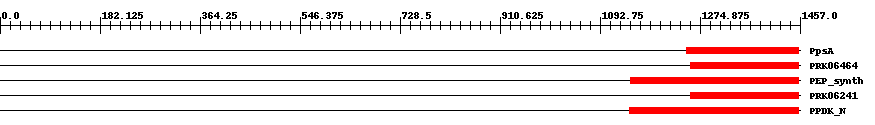 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0574 | PpsA | 2.0e-9 | 1251 | 1454 | 207 | + Phosphoenolpyruvate synthase/pyruvate phosphate dikinase [Carbohydrate transport and metabolism] | ||
| PRK06464 | PRK06464 | 6.0e-13 | 1257 | 1454 | 213 | + phosphoenolpyruvate synthase; Validated | ||
| TIGR01418 | PEP_synth | 6.0e-15 | 1148 | 1455 | 347 | + phosphoenolpyruvate synthase. Also called pyruvate,water dikinase and PEP synthase. The member from Methanococcus jannaschii contains a large intein. This enzyme generates phosphoenolpyruvate (PEP) from pyruvate, hydrolyzing ATP to AMP and releasing inorganic phosphate in the process. The enzyme shows extensive homology to other enzymes that use PEP as substrate or product. This enzyme may provide PEP for gluconeogenesis, for PTS-type carbohydrate transport systems, or for other processes [Energy metabolism, Glycolysis/gluconeogenesis]. | ||
| PRK06241 | PRK06241 | 6.0e-15 | 1257 | 1454 | 199 | + phosphoenolpyruvate synthase; Validated | ||
| pfam01326 | PPDK_N | 1.0e-33 | 1146 | 1455 | 335 | + Pyruvate phosphate dikinase, PEP/pyruvate binding domain. This enzyme catalyzes the reversible conversion of ATP to AMP, pyrophosphate and phosphoenolpyruvate (PEP). | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005524 | ATP binding |
| GO:0016301 | kinase activity |
| GO:0016310 | phosphorylation |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK11735.1 | 0 | 1 | 1456 | 5 | 1464 | starch associated protein R1 [Solanum tuberosum] |
| Swiss-Prot | Q8LPT9 | 0 | 1 | 1456 | 5 | 1475 | GWD1_CITRE RecName: Full=Alpha-glucan water dikinase, chloroplastic; AltName: Full=Starch-related protein R1; Flags: Precursor |
| RefSeq | XP_002270485.1 | 0 | 1 | 1456 | 5 | 1470 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002315679.1 | 0 | 5 | 1456 | 13 | 1477 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002527902.1 | 0 | 1 | 1456 | 1 | 1469 | alpha-glucan water dikinase, chloroplast precursor, putative [Ricinus communis] |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO777523 | 898 | 560 | 1457 | 0 |
| HO794398 | 752 | 706 | 1457 | 0 |
| HO794398 | 142 | 565 | 706 | 0 |
| HO779957 | 353 | 188 | 540 | 0 |
| HO779957 | 60 | 600 | 659 | 0 |
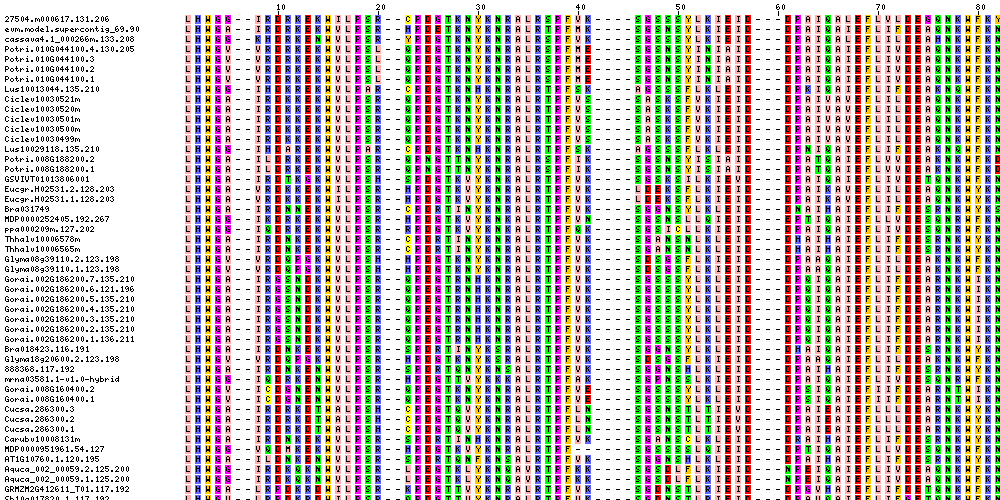
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |