

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s299_60V6.1 |
| Family | CBM22 |
| Protein Properties | Length: 1074 Molecular Weight: 119342 Isoelectric Point: 6.7199 |
| Chromosome | Chromosome/Scaffold: 299 Start: 385644 End: 391655 |
| Description | glycosyl hydrolase family 10 protein / carbohydrate-binding domain-containing protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM22 | 357 | 491 | 2.9e-21 |
| IVRNATFSQGLRNWKLRGSRGFVCNCLESPEVLPLEGKSFAVATQRIATWSGIEQTITDRIDLETIYEVAAAVRISGSCLKATVKATLYLREADNSERYI TMGTVEANTTEWKLLEGRMILLKAPSYVSVYLEGP | |||
| CBM22 | 6 | 142 | 5.5e-23 |
| NLIVNPTFIWGSDHGWQPICCSMSISDRLPQCGGPPSGHRFYCIAHNRTQVWQGIAQDISSRLKVGMEYKVEACVSISGPVELADVRVTLKLEHAEGQVT YTTLASGPVSKREWTFLNGNMEVSKGPIKALVYLEGP | |||
| CBM22 | 177 | 313 | 2.3e-25 |
| NVLVNPTFDTGVQGWSGICCKVAHSGMHGWKGIYGPRGKPFALAINRTEAWQGMEQDITTLVQSNETYLVTAVIRTAGKPHQGANVLATVRLDYGGSNPR YVSVGRVKASSTEWATLEGRLHLDTHPRKVVFYLEGP | |||
| CBM22 | 526 | 668 | 1e-30 |
| NIIENHDLQHGSKLWFGQGSARLSVASGAPTVVPPAAALSLPCSTSLSGNYLIAANRTQGWEGPAQNITDKLQLFVVYQVSAWVRVGKCRGKTGQKVNVA LSVDGSWMTGGEVEVDEYSWKEIMGSFRLEKKPNFAMVYAQGP | |||
| GH10 | 728 | 1011 | 0 |
| TSRSFPLGSCINRWSLNNNSYKQFFLQNFNWAVFENELKWNWIEPQRGTINYEDADEMVNFCSEHRIPMRGHCIFWEDESCCQDWLKTLSPLELKDALQN RAADLLRRYKGKFQHYDVNNEMLHGCFFRDRLNPDILPYVYKLAHQLEPEAVLFVNDYHVEDGVDANSAPEKYVKHIEWLRKEGAPIGAIGLQGHLDTPI GSIICNSLDKMSSVGLPLWMTEIDIAAANEHIRADDLEVVMRETFAHPSVEGIMLWGFWEGAMSRENGHLVDSDKRVNAAGERL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1074 Download |
| MAGSMNLIVN PTFIWGSDHG WQPICCSMSI SDRLPQCGGP PSGHRFYCIA HNRTQVWQGI 60 AQDISSRLKV GMEYKVEACV SISGPVELAD VRVTLKLEHA EGQVTYTTLA SGPVSKREWT 120 FLNGNMEVSK GPIKALVYLE GPPPGIDILA SCFSIALSKP ELQCSNKEPN EVNIHGNVLV 180 NPTFDTGVQG WSGICCKVAH SGMHGWKGIY GPRGKPFALA INRTEAWQGM EQDITTLVQS 240 NETYLVTAVI RTAGKPHQGA NVLATVRLDY GGSNPRYVSV GRVKASSTEW ATLEGRLHLD 300 THPRKVVFYL EGPPSGVDLL VSTVIIKHAN PPECEGSPLK RKRTEANSTR TGISDSIVRN 360 ATFSQGLRNW KLRGSRGFVC NCLESPEVLP LEGKSFAVAT QRIATWSGIE QTITDRIDLE 420 TIYEVAAAVR ISGSCLKATV KATLYLREAD NSERYITMGT VEANTTEWKL LEGRMILLKA 480 PSYVSVYLEG PPPETDILVD SFYIRPASKP EPPNPPIIKD PKYGVNIIEN HDLQHGSKLW 540 FGQGSARLSV ASGAPTVVPP AAALSLPCST SLSGNYLIAA NRTQGWEGPA QNITDKLQLF 600 VVYQVSAWVR VGKCRGKTGQ KVNVALSVDG SWMTGGEVEV DEYSWKEIMG SFRLEKKPNF 660 AMVYAQGPEP GIELMLAGLQ IFAVDRSARI PILKAQADKI RKRDVTIKLK PRNNRHIPPG 720 VCVRIEQTSR SFPLGSCINR WSLNNNSYKQ FFLQNFNWAV FENELKWNWI EPQRGTINYE 780 DADEMVNFCS EHRIPMRGHC IFWEDESCCQ DWLKTLSPLE LKDALQNRAA DLLRRYKGKF 840 QHYDVNNEML HGCFFRDRLN PDILPYVYKL AHQLEPEAVL FVNDYHVEDG VDANSAPEKY 900 VKHIEWLRKE GAPIGAIGLQ GHLDTPIGSI ICNSLDKMSS VGLPLWMTEI DIAAANEHIR 960 ADDLEVVMRE TFAHPSVEGI MLWGFWEGAM SRENGHLVDS DKRVNAAGER LINLREEWTT 1020 RLHGQMEESG QFSFRGYHGG YKAFVEYGEL GEIAVDFEVP KGDSPLVIEL SLS* 1080 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
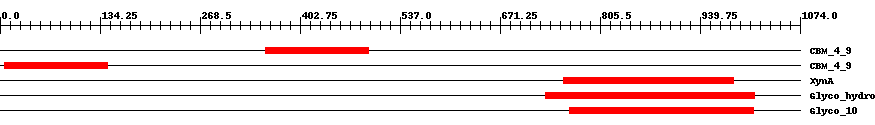 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam02018 | CBM_4_9 | 9.0e-13 | 357 | 495 | 141 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| pfam02018 | CBM_4_9 | 2.0e-13 | 6 | 144 | 140 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| COG3693 | XynA | 8.0e-35 | 756 | 985 | 252 | + Beta-1,4-xylanase [Carbohydrate transport and metabolism] | ||
| pfam00331 | Glyco_hydro_10 | 3.0e-70 | 732 | 1013 | 302 | + Glycosyl hydrolase family 10. | ||
| smart00633 | Glyco_10 | 3.0e-72 | 765 | 1011 | 268 | + Glycosyl hydrolase family 10. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| GO:0016798 | hydrolase activity, acting on glycosyl bonds |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_001770128.1 | 5e-32 | 4 | 326 | 155 | 482 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001770128.1 | 0 | 41 | 504 | 14 | 482 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001770128.1 | 0 | 200 | 1072 | 2 | 873 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001781282.1 | 0 | 41 | 504 | 10 | 479 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001781282.1 | 0 | 203 | 1073 | 1 | 871 | predicted protein [Physcomitrella patens subsp. patens] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1xyz_B | 8e-29 | 720 | 985 | 25 | 307 | A Chain A, A Common Protein Fold And Similar Active Site In Two Distinct Families Of Beta-Glycanases |
| PDB | 1xyz_A | 8e-29 | 720 | 985 | 25 | 307 | A Chain A, A Common Protein Fold And Similar Active Site In Two Distinct Families Of Beta-Glycanases |
| PDB | 1i1x_A | 3e-27 | 732 | 984 | 18 | 268 | A Chain A, A Common Protein Fold And Similar Active Site In Two Distinct Families Of Beta-Glycanases |
| PDB | 1i1w_A | 3e-27 | 732 | 984 | 18 | 268 | A Chain A, 0.89a Ultra High Resolution Structure Of A Thermostable Xylanase From Thermoascus Aurantiacus |
| PDB | 2bnj_A | 8e-27 | 732 | 984 | 18 | 268 | A Chain A, The Xylanase Ta From Thermoascus Aurantiacus Utilizes Arabinose Decorations Of Xylan As Significant Substrate Specificity Determinants |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| (1,4)-β-xylan degradation | 3.2.1.8-RXN | EC-3.2.1.8 | endo-1,4-β-xylanase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| BY972437 | 221 | 483 | 703 | 0 |
| FC330628 | 245 | 523 | 767 | 0 |
| BY972437 | 211 | 306 | 504 | 0.00000000000002 |
| BY972437 | 214 | 137 | 330 | 0.000000007 |
| BY972437 | 110 | 47 | 156 | 0.000004 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |