

| Basic Information | |
|---|---|
| Species | Mimulus guttatus |
| Cazyme ID | mgv1a006836m |
| Family | GT1 |
| Protein Properties | Length: 431 Molecular Weight: 48036.7 Isoelectric Point: 6.8435 |
| Chromosome | Chromosome/Scaffold: 122 Start: 606175 End: 609092 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
| Family | Start | End | Evalue |
| GT1 | 250 | 410 | 1.1e-32 |
| FWEEDKNCVDWLDEQEAGSVLYVSFGSWVSPIGEGKVEGLALTLEALGRPFVWVLGHAWRRGLPYGYADRVEVLKHPAVGCYLTHCGWNSTVEAVQCRKP LLCYPIAGDQFLNCAYIVDAWRIGVKIEGFGVEEVENGIREVVEDEKMRWRIERLNERLFG | |||
| Full Sequence |
|---|
| Protein Sequence Length: 431 Download |
| MKMKKKIIIL IPYPAQGHVT PMLKLASLLS NLGFRPVVVT PEFIHRRISP QINPRDGILC 60 VPISDGLDPA EPRDFFSIQR AMEETMPSNL EDIVRGMIHG GGGVACLVVD LLASWAVGVA 120 RQCGVEAVGF WPAMHATYRL VSAIPDLIRT GVISENGCPT NPKSRICLSP NEPTLAANEL 180 PWLIGSSSAR VSRFKFWTRT LERAKTLRYL LTNTFPNELD SKTQKTNMFP RVLKMGPLIM 240 QANTISTASF WEEDKNCVDW LDEQEAGSVL YVSFGSWVSP IGEGKVEGLA LTLEALGRPF 300 VWVLGHAWRR GLPYGYADRV EVLKHPAVGC YLTHCGWNST VEAVQCRKPL LCYPIAGDQF 360 LNCAYIVDAW RIGVKIEGFG VEEVENGIRE VVEDEKMRWR IERLNERLFG KEESSKAMAN 420 LSLFMEDLGE * |
| Functional Domains Download unfiltered results here | ||||||
|---|---|---|---|---|---|---|
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description |
| PLN02410 | PLN02410 | 1.0e-39 | 12 | 408 | 426 | + UDP-glucoronosyl/UDP-glucosyl transferase family protein |
| PLN02555 | PLN02555 | 1.0e-40 | 13 | 429 | 488 | + limonoid glucosyltransferase |
| PLN02173 | PLN02173 | 6.0e-43 | 12 | 406 | 443 | + UDP-glucosyl transferase family protein |
| PLN02448 | PLN02448 | 2.0e-43 | 12 | 430 | 469 | + UDP-glycosyltransferase family protein |
| PLN02562 | PLN02562 | 0 | 12 | 428 | 437 | + UDP-glycosyltransferase |
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAD44605.1 | 0 | 4 | 429 | 5 | 461 | hypothetical protein [Arabidopsis thaliana] |
| EMBL | CBI27396.1 | 0 | 3 | 412 | 4 | 429 | unnamed protein product [Vitis vinifera] |
| RefSeq | NP_188864.1 | 0 | 4 | 429 | 5 | 461 | UDP-glucoronosyl/UDP-glucosyl transferase family protein [Arabidopsis thaliana] |
| RefSeq | XP_002266304.1 | 0 | 3 | 428 | 4 | 445 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002521042.1 | 0 | 1 | 428 | 3 | 448 | UDP-glucosyltransferase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2pq6_A | 0 | 4 | 428 | 7 | 477 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 3hbj_A | 3e-30 | 53 | 419 | 73 | 442 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 3hbf_A | 3e-30 | 53 | 419 | 73 | 442 | A Chain A, Structure Of Ugt78g1 Complexed With Myricetin And Udp |
| PDB | 2vg8_A | 5e-26 | 4 | 423 | 5 | 465 | A Chain A, Structure Of Ugt78g1 Complexed With Myricetin And Udp |
| PDB | 2vch_A | 5e-26 | 4 | 423 | 5 | 465 | A Chain A, Structure Of Ugt78g1 Complexed With Myricetin And Udp |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
| Hit | Length | Start | End | EValue |
| GR121503 | 225 | 12 | 236 | 0 |
| DY288730 | 314 | 12 | 316 | 0 |
| DY273540 | 285 | 12 | 288 | 0 |
| DV136609 | 257 | 129 | 366 | 0 |
| GR121503 | 28 | 215 | 242 | 0.01 |
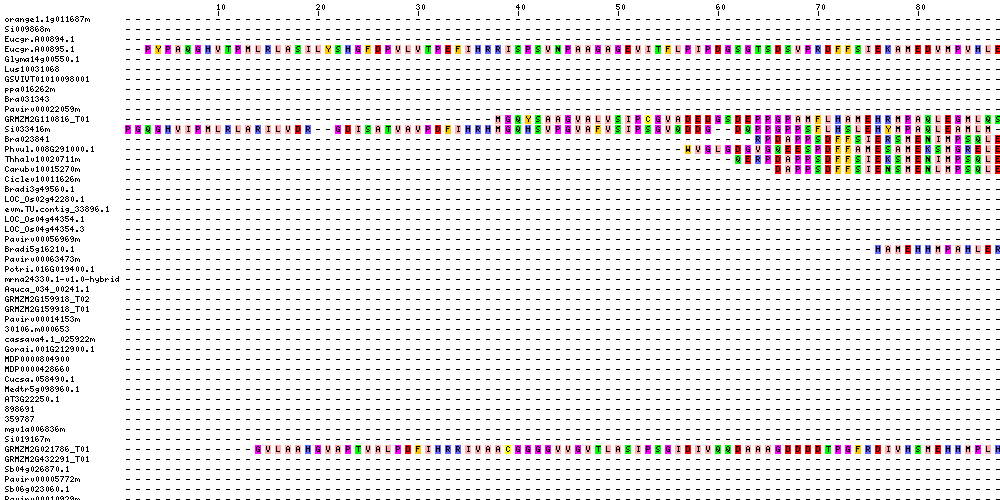
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |