

| Basic Information | |
|---|---|
| Species | Mimulus guttatus |
| Cazyme ID | mgv1a026906m |
| Family | GT1 |
| Protein Properties | Length: 472 Molecular Weight: 52879.5 Isoelectric Point: 5.477 |
| Chromosome | Chromosome/Scaffold: 19 Start: 105811 End: 107226 |
| Description | UDP-glucosyl transferase 73B3 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 240 | 442 | 2.2e-31 |
| GPILPAKLSSSANQSEIHNRCLEWLDKQEINSVIYVSFGTTTSLSDEQIKELAYGLEQSKVKFLWVLRDADKRDIFDGQVRRAELPDGFEERVEGVGMVV REWAPQPQILAHPSTGGFVSHCGWNSCIESITMGVPIAAWPMHSDQPNNSVLITEILKMGLLVRDWKQRTELVKASIIKNILKILMASKEGDELRKRAEE LAA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 472 Download |
| NQKEHTDTTT MEKPRVAVIM VPFPAQGHLN QLMQLSCLIS SYGIPVHYVG SAIHNRQAKV 60 RINGLNHEDV EKIHFRDIPT PSSVSPPPDP NSPDKPPTQL QPLWDSLSLR EPVASYLRDM 120 ANKFERVVVV HDSLMAVVVQ DVSSILNAES YVFNCISAFT KAVFTLDLLG KPLPIEHPKE 180 LPSFEGCITD EIRDFLALQF EPLSYRAGDL YNTCRLIEQD YMELIEKEEI VGSRKSWAIG 240 PILPAKLSSS ANQSEIHNRC LEWLDKQEIN SVIYVSFGTT TSLSDEQIKE LAYGLEQSKV 300 KFLWVLRDAD KRDIFDGQVR RAELPDGFEE RVEGVGMVVR EWAPQPQILA HPSTGGFVSH 360 CGWNSCIESI TMGVPIAAWP MHSDQPNNSV LITEILKMGL LVRDWKQRTE LVKASIIKNI 420 LKILMASKEG DELRKRAEEL AATVYQATQQ GGASKLELDS FISHITRGND I* 480 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN00164 | PLN00164 | 2.0e-46 | 207 | 454 | 260 | + glucosyltransferase; Provisional | ||
| PLN02863 | PLN02863 | 2.0e-46 | 208 | 467 | 267 | + UDP-glucoronosyl/UDP-glucosyl transferase family protein | ||
| PLN03007 | PLN03007 | 2.0e-50 | 208 | 466 | 271 | + UDP-glucosyltransferase family protein | ||
| PLN02448 | PLN02448 | 5.0e-51 | 237 | 469 | 239 | + UDP-glycosyltransferase family protein | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002278696.1 | 0 | 1 | 467 | 6 | 477 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002284381.1 | 0 | 1 | 467 | 3 | 473 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002308926.1 | 0 | 1 | 466 | 5 | 472 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002308931.1 | 0 | 1 | 466 | 5 | 472 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002322723.1 | 0 | 20 | 467 | 1 | 452 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2vg8_A | 0 | 14 | 454 | 7 | 456 | B Chain B, Structure Of A Cellulose Synthase - Cellulose Translocation Intermediate |
| PDB | 2vch_A | 0 | 14 | 454 | 7 | 456 | B Chain B, Structure Of A Cellulose Synthase - Cellulose Translocation Intermediate |
| PDB | 2vce_A | 0 | 14 | 454 | 7 | 456 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 2pq6_A | 1e-39 | 18 | 465 | 11 | 477 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2acw_B | 1e-36 | 237 | 469 | 242 | 465 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| GR164171 | 254 | 219 | 472 | 0 |
| GR173465 | 241 | 164 | 404 | 0 |
| GR156668 | 249 | 1 | 249 | 0 |
| GO956011 | 240 | 154 | 393 | 0 |
| EG631252 | 471 | 7 | 467 | 0 |
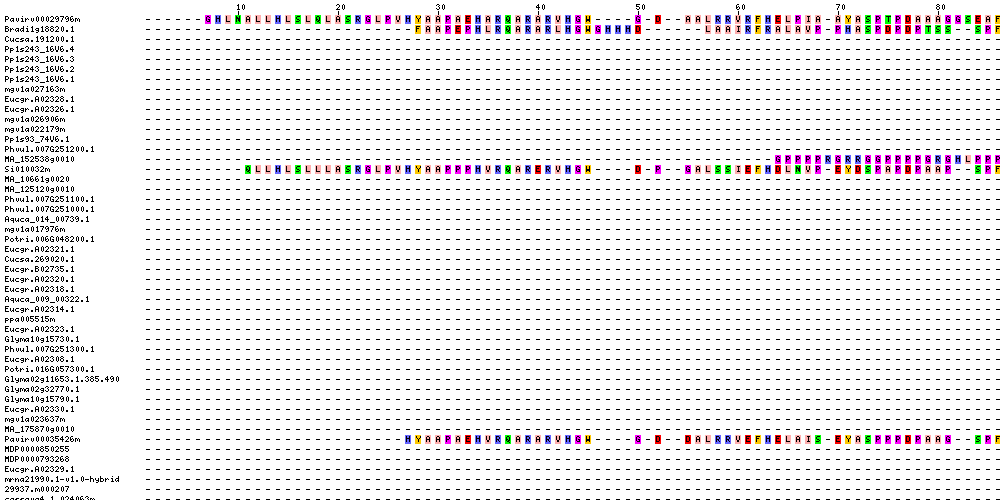
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |