

| Basic Information | |
|---|---|
| Species | Fragaria vesca |
| Cazyme ID | mrna04286.1-v1.0-hybrid |
| Family | GH13 |
| Protein Properties | Length: 1389 Molecular Weight: 155689 Isoelectric Point: 5.2595 |
| Chromosome | Chromosome/Scaffold: 6 Start: 32984988 End: 32996843 |
| Description | Transducin/WD40 repeat-like superfamily protein |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH13 | 1020 | 1304 | 1.8e-38 |
| LSKIGVTSVWFPPPTESVAPQGYMPSDLYNLNSAYGTEEELKYCIAEMHAHDLLALGDVVLNHRCAHKQSPNGVWNIFGGKLAWGPEAIVCDDPNFEGRG NPSSGDIFHAAPNIDHSKDFVRNDIKEWLNWLRSDIGFDGWRLDFVRGFSGSYVKEYIEASTPAFAIGEYWDSLAYENGNLCYNQDAHRQRIVNWINATG GSSSAFDVTTKGILHSALHNQYWRLIDPQGKPTGVMGWWPSRAVTFLENHDTGSTQGHWPFPREKLTQGYAYILTHPGTPTIFYD | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1389 Download |
| MEIDEGTEDR QVQVRFVTKL QDIPFKVPNT TIPASYSRLD LSKVVNALIQ TENPEWESEP 60 FDFLINGVLV RMSLGQFLLA KGISAEKTLE IEYIRAIVPR KEEEPCLHND WVSAVDGSSP 120 RFILTGCYDG LGRVWTASGT CTHILQGHSE PVTSVSVISA EGGESCRVAT ASKDRTLRLW 180 KFNTEEPTNN ALRTTAYKIL RGHGAAVQSV AAHTSGEMVC SGSWDSTIKL WQTDEPDIEV 240 DVSVKKRKKA EAVSSLVGHT QCVSSVKWPQ RDIIYSASWD HSIRRWDIES GKDILNIPCS 300 KALNCLDIGG EGSALVAGGG SDPILRIWDP RKPGTSAPVG QFSSHTSWIT ACKWHESSWF 360 HVLSASYDGK VMLWDMRTTW PLFVIDSHKD SKVLCVDWWK GDSVVSGGAD TKLCISSGVS 420 ALGLFPTSPC RIQVDIANMT ALYALGRGGL LSSELPLIEK RNYSSQNIGQ VNQEQAFFQA 480 WYADDSGVTD VVDNSDGFSS GRNDMLDVQE DKLMAVKRAL SEAQARQDAI EKERDQLLEE 540 LACAEAKQQE YVAAILHDKD MAVSELEAAK SLFDQKLQES VQEKFRLQDK LVLMKQDAVE 600 LAVQVERLAE IAFQQATSHI LEDSQLRVAA AETTAAEACY QIEKQIKDMT EGTILSIVEQ 660 SKNAIEKALD VAEKAGDHAT KAASAFSDSM SPLDELASVQ SKNIMLQGTV NDLESQLLLT 720 RSDVAKLKLE LEKAHAHTKL LEVRATDAEK ALVEFQDSSR KESLQREQEI MSLMEQMKKD 780 SSERNQAASG AFNVELQSIR DAIEAAKETV RSKDDAYLRR CEALQRSLKA SEATTKMWRQ 840 RAEIAESLLL KERQPADQEE DSIYVVNGGR IDLLTNDDSQ KWKLLSDGPR REIPQWMARR 900 ICTIRTNFPP RKIDVAEALS SEFRSLNLPK PEEVWSIALE KPKEGDTLVE HVFEKEILEK 960 KRKALERALQ RKSTQWQRTE EQTKLEPGTG TGREIVFQGF NWESWRRQWY LDLAPKAADL 1020 SKIGVTSVWF PPPTESVAPQ GYMPSDLYNL NSAYGTEEEL KYCIAEMHAH DLLALGDVVL 1080 NHRCAHKQSP NGVWNIFGGK LAWGPEAIVC DDPNFEGRGN PSSGDIFHAA PNIDHSKDFV 1140 RNDIKEWLNW LRSDIGFDGW RLDFVRGFSG SYVKEYIEAS TPAFAIGEYW DSLAYENGNL 1200 CYNQDAHRQR IVNWINATGG SSSAFDVTTK GILHSALHNQ YWRLIDPQGK PTGVMGWWPS 1260 RAVTFLENHD TGSTQGHWPF PREKLTQGYA YILTHPGTPT IFYDHLYDFG LHEILTELIE 1320 ARRRAGIHCR SAVKIYHANN EGYVAQVGDS LVMKLGHFDW NPSKENHLEG SWQKFVDQGA 1380 DYTVWLRQ* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
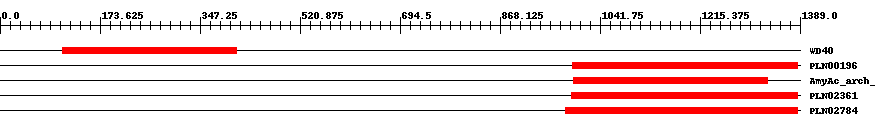 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| cd00200 | WD40 | 9.0e-49 | 108 | 410 | 308 | + WD40 domain, found in a number of eukaryotic proteins that cover a wide variety of functions including adaptor/regulatory modules in signal transduction, pre-mRNA processing and cytoskeleton assembly; typically contains a GH dipeptide 11-24 residues from its N-terminus and the WD dipeptide at its C-terminus and is 40 residues long, hence the name WD40; between GH and WD lies a conserved core; serves as a stable propeller-like platform to which proteins can bind either stably or reversibly; forms a propeller-like structure with several blades where each blade is composed of a four-stranded anti-parallel b-sheet; instances with few detectable copies are hypothesized to form larger structures by dimerization; each WD40 sequence repeat forms the first three strands of one blade and the last strand in the next blade; the last C-terminal WD40 repeat completes the blade structure of the first WD40 repeat to create the closed ring propeller-structure; residues on the top and bottom surface of the propeller are proposed to coordinate interactions with other proteins and/or small ligands; 7 copies of the repeat are present in this alignment. | ||
| PLN00196 | PLN00196 | 4.0e-140 | 994 | 1385 | 406 | + alpha-amylase; Provisional | ||
| cd11314 | AmyAc_arch_bac_plant_AmyA | 1.0e-162 | 995 | 1333 | 343 | + Alpha amylase catalytic domain found in archaeal, bacterial, and plant Alpha-amylases (also called 1,4-alpha-D-glucan-4-glucanohydrolase). AmyA (EC 3.2.1.1) catalyzes the hydrolysis of alpha-(1,4) glycosidic linkages of glycogen, starch, related polysaccharides, and some oligosaccharides. This group includes AmyA from bacteria, archaea, water fleas, and plants. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. | ||
| PLN02361 | PLN02361 | 8.0e-173 | 992 | 1385 | 399 | + alpha-amylase | ||
| PLN02784 | PLN02784 | 0 | 982 | 1385 | 408 | + alpha-amylase | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0004556 | alpha-amylase activity |
| GO:0005509 | calcium ion binding |
| GO:0005515 | protein binding |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3bsg_A | 0 | 993 | 1356 | 1 | 377 | A Chain A, Barley Alpha-Amylase Isozyme 1 (Amy1) H395a Mutant |
| PDB | 2qps_A | 0 | 993 | 1356 | 1 | 377 | A Chain A, "sugar Tongs" Mutant Y380a In Complex With Acarb |
| PDB | 1rpk_A | 0 | 993 | 1356 | 1 | 377 | A Chain A, "sugar Tongs" Mutant Y380a In Complex With Acarb |
| PDB | 1p6w_A | 0 | 993 | 1356 | 1 | 377 | A Chain A, "sugar Tongs" Mutant Y380a In Complex With Acarb |
| PDB | 1ht6_A | 0 | 993 | 1356 | 1 | 377 | A Chain A, Crystal Structure At 1.5a Resolution Of The Barley Alpha- Amylase Isozyme 1 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |