

| Basic Information | |
|---|---|
| Species | Prunus persica |
| Cazyme ID | ppa001089m |
| Family | CBM22 |
| Protein Properties | Length: 913 Molecular Weight: 100591 Isoelectric Point: 6.4395 |
| Chromosome | Chromosome/Scaffold: 1 Start: 35191178 End: 35195076 |
| Description | glycosyl hydrolase family 10 protein / carbohydrate-binding domain-containing protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM22 | 198 | 334 | 1.4e-32 |
| NIILNPKFDDGLNNWSGRGCKIVLHDSMGDGKIVPQTGKVFASATERTQSWNGIQQDVTGRLQRKLAYEATAVVRIFGNNVTSSDVRATLWVQSPNQREQ YIGIANVQATDKDWAQLQGKFLLNGSPSKVVVYLEGP | |||
| CBM22 | 27 | 163 | 7e-33 |
| NIILNHDFSGGLHSWHPNCCDGFVVSADSGHPEAKSAGNNYAVVNNRKECWQGLEQDITGRISPGSTYVVSACVGVSGPLQGSADVLATLKLEYQGSATN FLLIGRISVSNGRWETLDGKFSLSTMPDRVVFYLEGP | |||
| CBM22 | 369 | 510 | 2.99878e-43 |
| NIIENSNLSKGTNGWFPLGNCTLSVGTGSPHILPPMARDGLGPHEPLSGRYILVTKRTQTWMGPAQMIGDKLKLFLTYQVSAWVRIGAGATGPQNVNIAL GVDNQWVNGGQVEASDNRWHEIGGSFRIEKQPSKVMVYVQGP | |||
| GH10 | 567 | 845 | 0 |
| KQTKNSFPFGTCISRTNIDNEDFVDFFVKNFNWAVFGNELKWYWTEPQKGNFNYKDADELVDLCKSHNIDIRGHCIFWEVVDTVQQWIRSLSQNDLATAV QSRLTDLLTRYKGKFMHYDVNNEMLHGSFYQDKLGKDIRAKMFKSANQLDPSATLFVNDYHVEDGCDTRSSPERYIEHILDLQQQGAPVGGIGIQGHIDS PVGPIVCSALDKLGILGLPIWFTELDVSSVNEHVRADDLEVMLREGFANPAVEGIMMWGFWELFMSRQNSHLVNAEGDV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 913 Download |
| MENQKQTDNG ADHKEKLVNS SSSHATNIIL NHDFSGGLHS WHPNCCDGFV VSADSGHPEA 60 KSAGNNYAVV NNRKECWQGL EQDITGRISP GSTYVVSACV GVSGPLQGSA DVLATLKLEY 120 QGSATNFLLI GRISVSNGRW ETLDGKFSLS TMPDRVVFYL EGPSPGVDIL IKSVVISSSS 180 PKECQNGSSG NVNLGDENII LNPKFDDGLN NWSGRGCKIV LHDSMGDGKI VPQTGKVFAS 240 ATERTQSWNG IQQDVTGRLQ RKLAYEATAV VRIFGNNVTS SDVRATLWVQ SPNQREQYIG 300 IANVQATDKD WAQLQGKFLL NGSPSKVVVY LEGPPAGTDI LLNSFVVKHA ERVPPSPPPV 360 IENPAFGVNI IENSNLSKGT NGWFPLGNCT LSVGTGSPHI LPPMARDGLG PHEPLSGRYI 420 LVTKRTQTWM GPAQMIGDKL KLFLTYQVSA WVRIGAGATG PQNVNIALGV DNQWVNGGQV 480 EASDNRWHEI GGSFRIEKQP SKVMVYVQGP APGVDLMVAG VQIFPVDRQA RFKYLKRQTD 540 KIRKRDVVLK FSGLDSSSLL GCFVKVKQTK NSFPFGTCIS RTNIDNEDFV DFFVKNFNWA 600 VFGNELKWYW TEPQKGNFNY KDADELVDLC KSHNIDIRGH CIFWEVVDTV QQWIRSLSQN 660 DLATAVQSRL TDLLTRYKGK FMHYDVNNEM LHGSFYQDKL GKDIRAKMFK SANQLDPSAT 720 LFVNDYHVED GCDTRSSPER YIEHILDLQQ QGAPVGGIGI QGHIDSPVGP IVCSALDKLG 780 ILGLPIWFTE LDVSSVNEHV RADDLEVMLR EGFANPAVEG IMMWGFWELF MSRQNSHLVN 840 AEGDVNEAGK RYLELKKEWL SQAHGHIDEQ GEFIFRGFQG TYNIEIATAP KKLVKTFVVG 900 QGESPVEVPI AL* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
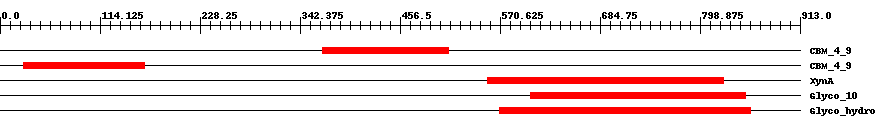 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam02018 | CBM_4_9 | 2.0e-15 | 368 | 512 | 153 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| pfam02018 | CBM_4_9 | 9.0e-19 | 27 | 165 | 140 | + Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. | ||
| COG3693 | XynA | 5.0e-33 | 556 | 826 | 294 | + Beta-1,4-xylanase [Carbohydrate transport and metabolism] | ||
| smart00633 | Glyco_10 | 2.0e-78 | 606 | 851 | 267 | + Glycosyl hydrolase family 10. | ||
| pfam00331 | Glyco_hydro_10 | 1.0e-79 | 570 | 856 | 307 | + Glycosyl hydrolase family 10. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| GO:0016798 | hydrolase activity, acting on glycosyl bonds |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAX33301.1 | 0 | 1 | 912 | 1 | 915 | putative endo-1,4-beta-xylanase [Populus tremula x Populus tremuloides] |
| RefSeq | XP_002283550.1 | 0 | 13 | 912 | 80 | 981 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002301133.1 | 0 | 15 | 912 | 152 | 1049 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 27 | 518 | 2 | 479 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002301133.1 | 0 | 197 | 523 | 1 | 314 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1nq6_A | 2e-28 | 575 | 847 | 15 | 291 | A Chain A, Crystal Structure Of The Catalytic Domain Of Xylanase A From Streptomyces Halstedii Jm8 |
| PDB | 1i1x_A | 5e-28 | 575 | 825 | 18 | 268 | A Chain A, Crystal Structure Of The Catalytic Domain Of Xylanase A From Streptomyces Halstedii Jm8 |
| PDB | 1i1w_A | 5e-28 | 575 | 825 | 18 | 268 | A Chain A, 0.89a Ultra High Resolution Structure Of A Thermostable Xylanase From Thermoascus Aurantiacus |
| PDB | 2bnj_A | 9e-28 | 575 | 825 | 18 | 268 | A Chain A, The Xylanase Ta From Thermoascus Aurantiacus Utilizes Arabinose Decorations Of Xylan As Significant Substrate Specificity Determinants |
| PDB | 1ta3_B | 2e-27 | 556 | 825 | 2 | 269 | B Chain B, Crystal Structure Of Xylanase (Gh10) In Complex With Inhibitor (Xip) |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EB442796 | 298 | 396 | 693 | 0 |
| EX570690 | 295 | 619 | 913 | 0 |
| GO850973 | 328 | 494 | 821 | 0 |
| EB442796 | 130 | 223 | 352 | 0.000003 |
| EB442796 | 100 | 67 | 166 | 0.0002 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |