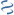MultiGeneBlast hits

Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
AP021873 : Serratia marcescens ATCC 274 DNA Total score: 1.0 Cumulative Blast bit score: 275
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
SMATCC274_08870
beta-glucoside operon transcriptional antiterminator
Accession: BBO61623
Location: 984528-985376
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 1e-87
NCBI BlastP on this gene
Accession: BBO61623
Location: 984528-985376
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 1e-87
NCBI BlastP on this gene
SMATCC274_08860
twitching motility protein PilT
Accession: BBO61622
Location: 984145-984531
NCBI BlastP on this gene
Accession: BBO61622
Location: 984145-984531
NCBI BlastP on this gene
SMATCC274_08850
SMATCC274_08840
SMATCC274_08830
recombination-associated protein RdgC
Accession: BBO61619
Location: 981769-982680
NCBI BlastP on this gene
Accession: BBO61619
Location: 981769-982680
NCBI BlastP on this gene
rdgC
SMATCC274_08810
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
AP019009 : Serratia marcescens AS-1 DNA Total score: 1.0 Cumulative Blast bit score: 275
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
beta-glucoside operon transcriptional antiterminator
Accession: BBG68099
Location: 1018724-1019572
BlastP hit with bglK
Percentage identity: 49 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 4e-88
NCBI BlastP on this gene
Accession: BBG68099
Location: 1018724-1019572
BlastP hit with bglK
Percentage identity: 49 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 4e-88
NCBI BlastP on this gene
SERAS_09020
twitching motility protein PilT
Accession: BBG68098
Location: 1018341-1018727
NCBI BlastP on this gene
Accession: BBG68098
Location: 1018341-1018727
NCBI BlastP on this gene
SERAS_09010
SERAS_09000
SERAS_08990
recombination-associated protein RdgC
Accession: BBG68095
Location: 1015965-1016876
NCBI BlastP on this gene
Accession: BBG68095
Location: 1015965-1016876
NCBI BlastP on this gene
rdgC
SERAS_08970
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP047605 : Serratia sp. NGAS9 chromosome Total score: 1.0 Cumulative Blast bit score: 274
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QHI76891
Location: 977103-977951
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: QHI76891
Location: 977103-977951
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
GUC32_04605
GUC32_04600
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QHI76889
Location: 976490-976723
NCBI BlastP on this gene
Accession: QHI76889
Location: 976490-976723
NCBI BlastP on this gene
GUC32_04595
GUC32_04590
recombination-associated protein RdgC
Accession: QHI76887
Location: 974345-975256
NCBI BlastP on this gene
Accession: QHI76887
Location: 974345-975256
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QHI76886
Location: 973989-974279
NCBI BlastP on this gene
Accession: QHI76886
Location: 973989-974279
NCBI BlastP on this gene
ppnP
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP041124 : Serratia marcescens strain WVU-003 chromosome Total score: 1.0 Cumulative Blast bit score: 274
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QDI22246
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: QDI22246
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
FBF90_04720
type II toxin-antitoxin system VapC family toxin
Accession: QDI22245
Location: 1011968-1012354
NCBI BlastP on this gene
Accession: QDI22245
Location: 1011968-1012354
NCBI BlastP on this gene
FBF90_04715
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QDI22244
Location: 1011738-1011971
NCBI BlastP on this gene
Accession: QDI22244
Location: 1011738-1011971
NCBI BlastP on this gene
FBF90_04710
FBF90_04705
recombination-associated protein RdgC
Accession: QDI22242
Location: 1009593-1010504
NCBI BlastP on this gene
Accession: QDI22242
Location: 1009593-1010504
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QDI22241
Location: 1009237-1009527
NCBI BlastP on this gene
Accession: QDI22241
Location: 1009237-1009527
NCBI BlastP on this gene
ppnP
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP041122 : Serratia marcescens strain WVU-001 chromosome Total score: 1.0 Cumulative Blast bit score: 274
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QDI12503
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: QDI12503
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
FBF84_04720
type II toxin-antitoxin system VapC family toxin
Accession: QDI12502
Location: 1011968-1012354
NCBI BlastP on this gene
Accession: QDI12502
Location: 1011968-1012354
NCBI BlastP on this gene
FBF84_04715
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QDI12501
Location: 1011738-1011971
NCBI BlastP on this gene
Accession: QDI12501
Location: 1011738-1011971
NCBI BlastP on this gene
FBF84_04710
FBF84_04705
recombination-associated protein RdgC
Accession: QDI12499
Location: 1009593-1010504
NCBI BlastP on this gene
Accession: QDI12499
Location: 1009593-1010504
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QDI12498
Location: 1009237-1009527
NCBI BlastP on this gene
Accession: QDI12498
Location: 1009237-1009527
NCBI BlastP on this gene
ppnP
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP018930 : Serratia marcescens strain UMH12 chromosome Total score: 1.0 Cumulative Blast bit score: 274
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
BVG84_01565
transcription antiterminator BglG
Accession: ASM29766
Location: 322310-323158
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: ASM29766
Location: 322310-323158
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
BVG84_01560
twitching motility protein PilT
Accession: ASM29765
Location: 321927-322313
NCBI BlastP on this gene
Accession: ASM29765
Location: 321927-322313
NCBI BlastP on this gene
BVG84_01555
BVG84_01550
BVG84_01545
recombination-associated protein RdgC
Accession: ASM29762
Location: 319552-320463
NCBI BlastP on this gene
Accession: ASM29762
Location: 319552-320463
NCBI BlastP on this gene
BVG84_01540
BVG84_01535
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP018915 : Serratia marcescens strain UMH1 chromosome Total score: 1.0 Cumulative Blast bit score: 274
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
BVG95_01540
transcription antiterminator BglG
Accession: ASL81665
Location: 319631-320479
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: ASL81665
Location: 319631-320479
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
BVG95_01535
twitching motility protein PilT
Accession: ASL81664
Location: 319248-319634
NCBI BlastP on this gene
Accession: ASL81664
Location: 319248-319634
NCBI BlastP on this gene
BVG95_01530
BVG95_01525
BVG95_01520
recombination-associated protein RdgC
Accession: ASL81661
Location: 316873-317784
NCBI BlastP on this gene
Accession: ASL81661
Location: 316873-317784
NCBI BlastP on this gene
BVG95_01515
BVG95_01510
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
LT906479 : Serratia ficaria strain NCTC12148 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 273
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
serine/threonine protein kinase
Accession: SNV90830
Location: 1140607-1141617
NCBI BlastP on this gene
Accession: SNV90830
Location: 1140607-1141617
NCBI BlastP on this gene
SAMEA4384070_01073
4-aminobutyrate aminotransferase GabT
Accession: SNV90820
Location: 1139047-1140354
NCBI BlastP on this gene
Accession: SNV90820
Location: 1139047-1140354
NCBI BlastP on this gene
gabT
Low-affinity putrescine importer PlaP
Accession: SNV90812
Location: 1137617-1138984
NCBI BlastP on this gene
Accession: SNV90812
Location: 1137617-1138984
NCBI BlastP on this gene
plaP_1
Transcription antiterminator LicT
Accession: SNV90802
Location: 1136588-1137430
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 273
Sequence coverage: 100 %
E-value: 3e-87
NCBI BlastP on this gene
Accession: SNV90802
Location: 1136588-1137430
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 273
Sequence coverage: 100 %
E-value: 3e-87
NCBI BlastP on this gene
licT_1
mak
Transcriptional regulatory protein OmpR
Accession: SNV90786
Location: 1134646-1135380
NCBI BlastP on this gene
Accession: SNV90786
Location: 1134646-1135380
NCBI BlastP on this gene
ompR_1
ABC-type sugar transport system, auxiliary component
Accession: SNV90777
Location: 1133904-1134590
NCBI BlastP on this gene
Accession: SNV90777
Location: 1133904-1134590
NCBI BlastP on this gene
SAMEA4384070_01067
D-ribose-binding periplasmic protein precursor
Accession: SNV90768
Location: 1132904-1133845
NCBI BlastP on this gene
Accession: SNV90768
Location: 1132904-1133845
NCBI BlastP on this gene
rbsB_2
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP047391 : Serratia marcescens strain 1602 chromosome Total score: 1.0 Cumulative Blast bit score: 272
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QHE55762
Location: 983893-984741
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: QHE55762
Location: 983893-984741
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
GRX59_04605
GRX59_04600
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QHE55760
Location: 983280-983513
NCBI BlastP on this gene
Accession: QHE55760
Location: 983280-983513
NCBI BlastP on this gene
GRX59_04595
GRX59_04590
recombination-associated protein RdgC
Accession: QHE55758
Location: 981135-982046
NCBI BlastP on this gene
Accession: QHE55758
Location: 981135-982046
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QHE55757
Location: 980779-981069
NCBI BlastP on this gene
Accession: QHE55757
Location: 980779-981069
NCBI BlastP on this gene
ppnP
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP029746 : Serratia marcescens strain AR_0122 Total score: 1.0 Cumulative Blast bit score: 272
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AM369_17260
PRD domain-containing protein
Accession: AWS59912
Location: 3659040-3659888
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWS59912
Location: 3659040-3659888
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM369_17255
AM369_17250
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWS59910
Location: 3658427-3658660
NCBI BlastP on this gene
Accession: AWS59910
Location: 3658427-3658660
NCBI BlastP on this gene
AM369_17245
AM369_17240
recombination-associated protein RdgC
Accession: AWS59908
Location: 3656282-3657193
NCBI BlastP on this gene
Accession: AWS59908
Location: 3656282-3657193
NCBI BlastP on this gene
AM369_17235
pyrimidine/purine nucleoside phosphorylase
Accession: AWS59907
Location: 3655926-3656216
NCBI BlastP on this gene
Accession: AWS59907
Location: 3655926-3656216
NCBI BlastP on this gene
AM369_17230
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP029715 : Serratia marcescens strain AR_0131 chromosome Total score: 1.0 Cumulative Blast bit score: 272
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
PRD domain-containing protein
Accession: AWS71293
Location: 5081706-5082554
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWS71293
Location: 5081706-5082554
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM378_24130
AM378_24125
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWS71291
Location: 5081093-5081326
NCBI BlastP on this gene
Accession: AWS71291
Location: 5081093-5081326
NCBI BlastP on this gene
AM378_24120
AM378_24115
recombination-associated protein RdgC
Accession: AWS71289
Location: 5078948-5079859
NCBI BlastP on this gene
Accession: AWS71289
Location: 5078948-5079859
NCBI BlastP on this gene
AM378_24110
pyrimidine/purine nucleoside phosphorylase
Accession: AWS71288
Location: 5078592-5078882
NCBI BlastP on this gene
Accession: AWS71288
Location: 5078592-5078882
NCBI BlastP on this gene
AM378_24105
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP028949 : Serratia marcescens strain AR_0121 chromosome Total score: 1.0 Cumulative Blast bit score: 272
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AM368_06645
PRD domain-containing protein
Accession: AWC69937
Location: 1377712-1378560
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWC69937
Location: 1377712-1378560
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM368_06650
AM368_06655
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWC69939
Location: 1378940-1379173
NCBI BlastP on this gene
Accession: AWC69939
Location: 1378940-1379173
NCBI BlastP on this gene
AM368_06660
AM368_06665
recombination-associated protein RdgC
Accession: AWC69941
Location: 1380407-1381318
NCBI BlastP on this gene
Accession: AWC69941
Location: 1380407-1381318
NCBI BlastP on this gene
AM368_06670
pyrimidine/purine nucleoside phosphorylase
Accession: AWC69942
Location: 1381384-1381674
NCBI BlastP on this gene
Accession: AWC69942
Location: 1381384-1381674
NCBI BlastP on this gene
AM368_06675
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP028948 : Serratia marcescens strain AR_0123 chromosome Total score: 1.0 Cumulative Blast bit score: 272
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AM370_02385
PRD domain-containing protein
Accession: AWC87888
Location: 477583-478431
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWC87888
Location: 477583-478431
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM370_02390
AM370_02395
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWC87890
Location: 478811-479044
NCBI BlastP on this gene
Accession: AWC87890
Location: 478811-479044
NCBI BlastP on this gene
AM370_02400
AM370_02405
recombination-associated protein RdgC
Accession: AWC87892
Location: 480278-481189
NCBI BlastP on this gene
Accession: AWC87892
Location: 480278-481189
NCBI BlastP on this gene
AM370_02410
pyrimidine/purine nucleoside phosphorylase
Accession: AWC87893
Location: 481255-481545
NCBI BlastP on this gene
Accession: AWC87893
Location: 481255-481545
NCBI BlastP on this gene
AM370_02415
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
LT883155 : Serratia grimesii isolate BXF1 genome assembly, chromosome. Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
Transcription antiterminator LicT
Accession: SMZ55352
Location: 1038009-1038851
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: SMZ55352
Location: 1038009-1038851
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 3e-86
NCBI BlastP on this gene
licT_1
mak
Transcriptional regulatory protein OmpR
Accession: SMZ55350
Location: 1036071-1036811
NCBI BlastP on this gene
Accession: SMZ55350
Location: 1036071-1036811
NCBI BlastP on this gene
ompR_1
ydjH_1
putative fructose-bisphosphate aldolase
Accession: SMZ55348
Location: 1034223-1035083
NCBI BlastP on this gene
Accession: SMZ55348
Location: 1034223-1035083
NCBI BlastP on this gene
fbaA_1
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP050171 : Serratia fonticola strain CPSE11 chromosome Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
HAP32_03519
transcription antiterminator BglG
Accession: QIP92998
Location: 3898814-3899671
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: QIP92998
Location: 3898814-3899671
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
HAP32_03518
HAP32_03517
DNA recombination-dependent growth factor C
Accession: QIP92996
Location: 3896565-3897476
NCBI BlastP on this gene
Accession: QIP92996
Location: 3896565-3897476
NCBI BlastP on this gene
HAP32_03516
HAP32_03515
HAP32_03514
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP040182 : Serratia fonticola strain MS5 chromosome Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QCR60083
Location: 1508365-1509222
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: QCR60083
Location: 1508365-1509222
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
FD644_06815
FD644_06810
recombination-associated protein RdgC
Accession: QCR60081
Location: 1506126-1507037
NCBI BlastP on this gene
Accession: QCR60081
Location: 1506126-1507037
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QCR60080
Location: 1505759-1506049
NCBI BlastP on this gene
Accession: QCR60080
Location: 1505759-1506049
NCBI BlastP on this gene
ppnP
DUF1177 domain-containing protein
Accession: QCR60079
Location: 1504645-1505574
NCBI BlastP on this gene
Accession: QCR60079
Location: 1504645-1505574
NCBI BlastP on this gene
FD644_06795
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP031316 : Serratia marcescens strain N4-5 chromosome Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
Exonuclease SbcC family protein
Accession: AXK22751
Location: 993288-996539
NCBI BlastP on this gene
Accession: AXK22751
Location: 993288-996539
NCBI BlastP on this gene
SmN45_0945
Transcription antiterminator LicT
Accession: AXK22750
Location: 992208-993056
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AXK22750
Location: 992208-993056
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 1e-86
NCBI BlastP on this gene
licT
SmN45_0943
Prevent-host-death family protein
Accession: AXK22748
Location: 991595-991828
NCBI BlastP on this gene
Accession: AXK22748
Location: 991595-991828
NCBI BlastP on this gene
SmN45_0942
SmN45_0941
Recombination-associated protein RdgC
Accession: AXK22746
Location: 989449-990360
NCBI BlastP on this gene
Accession: AXK22746
Location: 989449-990360
NCBI BlastP on this gene
rdgC
SmN45_0939
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP023956 : Serratia fonticola strain FDAARGOS_411 chromosome Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
CRN79_04260
transcription antiterminator BglG
Accession: ATM75107
Location: 960943-961800
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 2e-86
NCBI BlastP on this gene
Accession: ATM75107
Location: 960943-961800
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 2e-86
NCBI BlastP on this gene
CRN79_04265
CRN79_04270
recombination-associated protein RdgC
Accession: ATM75109
Location: 963009-963920
NCBI BlastP on this gene
Accession: ATM75109
Location: 963009-963920
NCBI BlastP on this gene
CRN79_04275
pyrimidine/purine nucleoside phosphorylase
Accession: ATM75110
Location: 963997-964287
NCBI BlastP on this gene
Accession: ATM75110
Location: 963997-964287
NCBI BlastP on this gene
CRN79_04280
CRN79_04285
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP014017 : Serratia liquefaciens strain FDAARGOS_125 chromosome Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AL485_07500
PRD domain-containing protein
Accession: AMG99026
Location: 1582508-1583350
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AMG99026
Location: 1582508-1583350
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AL485_07505
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP011254 : Serratia fonticola strain DSM 4576 Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: AKG70130
Location: 3169195-3172446
NCBI BlastP on this gene
Accession: AKG70130
Location: 3169195-3172446
NCBI BlastP on this gene
WN53_14050
transcription antiterminator BglG
Accession: AKG70129
Location: 3168131-3168988
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: AKG70129
Location: 3168131-3168988
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
WN53_14045
WN53_14040
recombination associated protein
Accession: AKG70127
Location: 3166021-3166932
NCBI BlastP on this gene
Accession: AKG70127
Location: 3166021-3166932
NCBI BlastP on this gene
WN53_14035
WN53_14030
WN53_14025
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP006252 : Serratia liquefaciens ATCC 27592 Total score: 1.0 Cumulative Blast bit score: 271
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: AGQ29717
Location: 959249-962500
NCBI BlastP on this gene
Accession: AGQ29717
Location: 959249-962500
NCBI BlastP on this gene
M495_04455
beta-1,4-xylanase
Accession: AGQ29716
Location: 958211-959077
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: AGQ29716
Location: 958211-959077
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 4e-86
NCBI BlastP on this gene
M495_04450
M495_04445
M495_04440
M495_04435
M495_04430
M495_04425
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
LS483469 : Serratia plymuthica strain NCTC12961 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC_2
sbcC_1
HTH-type transcriptional regulator gadW
Accession: SQI32688
Location: 1059059-1059520
NCBI BlastP on this gene
Accession: SQI32688
Location: 1059059-1059520
NCBI BlastP on this gene
gadW
Proline-specific permease ProY
Accession: SQI32685
Location: 1057421-1058788
NCBI BlastP on this gene
Accession: SQI32685
Location: 1057421-1058788
NCBI BlastP on this gene
proY_2
Transcription antiterminator LicT
Accession: SQI32683
Location: 1056304-1057146
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: SQI32683
Location: 1056304-1057146
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
licT_1
NCTC12961_01230
Antitoxin of toxin-antitoxin stability system
Accession: SQI32681
Location: 1055685-1055918
NCBI BlastP on this gene
Accession: SQI32681
Location: 1055685-1055918
NCBI BlastP on this gene
NCTC12961_01229
mak
Transcriptional regulatory protein OmpR
Accession: SQI32679
Location: 1053742-1054482
NCBI BlastP on this gene
Accession: SQI32679
Location: 1053742-1054482
NCBI BlastP on this gene
ompR_1
NCTC12961_01226
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP033893 : Serratia liquefaciens strain FG3 chromosome Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QDL35495
Location: 3682954-3683796
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: QDL35495
Location: 3682954-3683796
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
EGO53_18000
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP033162 : Serratia sp. P2ACOL2 chromosome Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: AYO40686
Location: 831433-832275
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: AYO40686
Location: 831433-832275
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
EBA31_04005
EBA31_04000
EBA31_03995
EBA31_03990
carbohydrate kinase family protein
Accession: AYO36504
Location: 828328-829341
NCBI BlastP on this gene
Accession: AYO36504
Location: 828328-829341
NCBI BlastP on this gene
EBA31_03985
ketose 1,6-bisphosphate aldolase
Accession: AYO36503
Location: 827468-828328
NCBI BlastP on this gene
Accession: AYO36503
Location: 827468-828328
NCBI BlastP on this gene
EBA31_03980
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP033055 : Serratia sp. 3ACOL1 chromosome Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
PRD domain-containing protein
Accession: AYM93815
Location: 5832446-5833303
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 270
Sequence coverage: 98 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AYM93815
Location: 5832446-5833303
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 270
Sequence coverage: 98 %
E-value: 8e-86
NCBI BlastP on this gene
D9980_26435
D9980_26430
recombination-associated protein RdgC
Accession: AYM93813
Location: 5830336-5831247
NCBI BlastP on this gene
Accession: AYM93813
Location: 5830336-5831247
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: AYM93812
Location: 5829969-5830259
NCBI BlastP on this gene
Accession: AYM93812
Location: 5829969-5830259
NCBI BlastP on this gene
D9980_26420
DUF1177 domain-containing protein
Accession: AYM93811
Location: 5828855-5829784
NCBI BlastP on this gene
Accession: AYM93811
Location: 5828855-5829784
NCBI BlastP on this gene
D9980_26415
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP012097 : Serratia plymuthica strain 3Re4-18 Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: ANJ97339
Location: 1092870-1096124
NCBI BlastP on this gene
Accession: ANJ97339
Location: 1092870-1096124
NCBI BlastP on this gene
ADP73_05040
ADP73_05035
ADP73_05030
transcription antiterminator BglG
Accession: ANJ97337
Location: 1089659-1090501
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: ANJ97337
Location: 1089659-1090501
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
ADP73_05025
ADP73_05020
ADP73_05015
ADP73_05010
ADP73_05005
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP012096 : Serratia plymuthica strain 3Rp8 Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: ANJ92596
Location: 1264435-1267689
NCBI BlastP on this gene
Accession: ANJ92596
Location: 1264435-1267689
NCBI BlastP on this gene
ADP72_06195
ADP72_06200
ADP72_06205
transcription antiterminator BglG
Accession: ANJ92598
Location: 1270058-1270900
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: ANJ92598
Location: 1270058-1270900
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
ADP72_06210
ADP72_06215
ADP72_06220
ADP72_06225
ADP72_06230
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP007439 : Serratia plymuthica strain V4 genome. Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: AHY05951
Location: 1071599-1074853
NCBI BlastP on this gene
Accession: AHY05951
Location: 1071599-1074853
NCBI BlastP on this gene
sch_05035
sch_05030
sch_05025
transcription antiterminator BglG
Accession: AHY05948
Location: 1068388-1069230
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AHY05948
Location: 1068388-1069230
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
sch_05020
sch_05015
sch_05010
sch_05005
sch_05000
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP006566 : Serratia plymuthica S13 Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: AGP43241
Location: 1039188-1042442
NCBI BlastP on this gene
Accession: AGP43241
Location: 1039188-1042442
NCBI BlastP on this gene
M621_04885
M621_04880
M621_04875
beta-1,4-xylanase
Accession: AGP43239
Location: 1035977-1036819
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AGP43239
Location: 1035977-1036819
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
M621_04870
M621_04865
M621_04860
M621_04855
M621_04850
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP006250 : Serratia plymuthica 4Rx13 Total score: 1.0 Cumulative Blast bit score: 270
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
transcriptional regulatory protein
Accession: AGO53887
Location: 972639-973100
NCBI BlastP on this gene
Accession: AGO53887
Location: 972639-973100
NCBI BlastP on this gene
SOD_c08900
SOD_c08890
beta-glucoside operon antiterminator
Accession: AGO53885
Location: 969887-970729
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AGO53885
Location: 969887-970729
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
arbG
mak
transcriptional regulatory protein OmpR
Accession: AGO53883
Location: 967947-968687
NCBI BlastP on this gene
Accession: AGO53883
Location: 967947-968687
NCBI BlastP on this gene
ompR1
ribokinase-like domain-containing protein
Accession: AGO53882
Location: 966959-967972
NCBI BlastP on this gene
Accession: AGO53882
Location: 966959-967972
NCBI BlastP on this gene
SOD_c08850
D-tagatose-1,6-bisphosphate aldolase subunit GatY
Accession: AGO53881
Location: 966099-966959
NCBI BlastP on this gene
Accession: AGO53881
Location: 966099-966959
NCBI BlastP on this gene
gatY1
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP048784 : Serratia liquefaciens strain S1 chromosome Total score: 1.0 Cumulative Blast bit score: 269
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
PRD domain-containing protein
Accession: QIC85739
Location: 997649-998491
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
Accession: QIC85739
Location: 997649-998491
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
F0336_04650
type II toxin-antitoxin system VapC family toxin
Accession: F0336_04645
Location: 997420-997562
NCBI BlastP on this gene
Accession: F0336_04645
Location: 997420-997562
NCBI BlastP on this gene
F0336_04645
mak
response regulator transcription factor
Accession: QIC85737
Location: 995466-996206
NCBI BlastP on this gene
Accession: QIC85737
Location: 995466-996206
NCBI BlastP on this gene
F0336_04635
carbohydrate kinase family protein
Accession: QIC85736
Location: 994478-995491
NCBI BlastP on this gene
Accession: QIC85736
Location: 994478-995491
NCBI BlastP on this gene
F0336_04630
ketose 1,6-bisphosphate aldolase
Accession: QIC85735
Location: 993618-994478
NCBI BlastP on this gene
Accession: QIC85735
Location: 993618-994478
NCBI BlastP on this gene
F0336_04625
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP011303 : Serratia liquefaciens strain HUMV-21 Total score: 1.0 Cumulative Blast bit score: 269
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: AKE11835
Location: 4016221-4019472
NCBI BlastP on this gene
Accession: AKE11835
Location: 4016221-4019472
NCBI BlastP on this gene
XJ20_18795
transcription antiterminator BglG
Accession: AKE13125
Location: 4019668-4020510
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
Accession: AKE13125
Location: 4019668-4020510
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
XJ20_18800
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP002775 : Serratia sp. AS13 Total score: 1.0 Cumulative Blast bit score: 269
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
SerAS13_0964
SerAS13_0962
transcriptional antiterminator, BglG
Accession: AEG26773
Location: 1060650-1061492
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
Accession: AEG26773
Location: 1060650-1061492
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
SerAS13_0961
SerAS13_0960
prevent-host-death family protein
Accession: AEG26771
Location: 1060031-1060264
NCBI BlastP on this gene
Accession: AEG26771
Location: 1060031-1060264
NCBI BlastP on this gene
SerAS13_0959
SerAS13_0958
two component transcriptional regulator, winged helix family
Accession: AEG26769
Location: 1058089-1058829
NCBI BlastP on this gene
Accession: AEG26769
Location: 1058089-1058829
NCBI BlastP on this gene
SerAS13_0957
SerAS13_0956
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP002774 : Serratia sp. AS12 Total score: 1.0 Cumulative Blast bit score: 269
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
SerAS12_0964
SerAS12_0962
transcriptional antiterminator, BglG
Accession: AEF49065
Location: 1060653-1061495
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
Accession: AEF49065
Location: 1060653-1061495
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
SerAS12_0961
SerAS12_0960
prevent-host-death family protein
Accession: AEF49063
Location: 1060034-1060267
NCBI BlastP on this gene
Accession: AEF49063
Location: 1060034-1060267
NCBI BlastP on this gene
SerAS12_0959
SerAS12_0958
two component transcriptional regulator, winged helix family
Accession: AEF49061
Location: 1058092-1058832
NCBI BlastP on this gene
Accession: AEF49061
Location: 1058092-1058832
NCBI BlastP on this gene
SerAS12_0957
SerAS12_0956
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP002773 : Serratia plymuthica AS9 Total score: 1.0 Cumulative Blast bit score: 269
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
SerAS9_0964
SerAS9_0962
transcriptional antiterminator, BglG
Accession: AEF44113
Location: 1060838-1061680
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
Accession: AEF44113
Location: 1060838-1061680
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
SerAS9_0961
SerAS9_0960
prevent-host-death family protein
Accession: AEF44111
Location: 1060219-1060452
NCBI BlastP on this gene
Accession: AEF44111
Location: 1060219-1060452
NCBI BlastP on this gene
SerAS9_0959
SerAS9_0958
two component transcriptional regulator, winged helix family
Accession: AEF44109
Location: 1058277-1059017
NCBI BlastP on this gene
Accession: AEF44109
Location: 1058277-1059017
NCBI BlastP on this gene
SerAS9_0957
SerAS9_0956
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
LR134478 : Serratia plymuthica strain NCTC8015 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 267
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
Low-affinity putrescine importer PlaP
Accession: VEI16987
Location: 1030649-1032016
NCBI BlastP on this gene
Accession: VEI16987
Location: 1030649-1032016
NCBI BlastP on this gene
plaP_1
Transcription antiterminator LicT
Accession: VEI16985
Location: 1029533-1030375
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
Accession: VEI16985
Location: 1029533-1030375
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
licT_1
NCTC8015_00990
Antitoxin of toxin-antitoxin stability system
Accession: VEI16981
Location: 1028914-1029147
NCBI BlastP on this gene
Accession: VEI16981
Location: 1028914-1029147
NCBI BlastP on this gene
NCTC8015_00989
mak
Transcriptional regulatory protein OmpR
Accession: VEI16977
Location: 1026971-1027711
NCBI BlastP on this gene
Accession: VEI16977
Location: 1026971-1027711
NCBI BlastP on this gene
ompR_1
rbsK_1
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
LR134151 : Serratia plymuthica strain NCTC8900 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 267
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC_1
Low-affinity putrescine importer PlaP
Accession: VEA63288
Location: 1019379-1020746
NCBI BlastP on this gene
Accession: VEA63288
Location: 1019379-1020746
NCBI BlastP on this gene
plaP_1
Transcription antiterminator LicT
Accession: VEA63286
Location: 1018263-1019105
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
Accession: VEA63286
Location: 1018263-1019105
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
licT_1
NCTC8900_01022
Antitoxin of toxin-antitoxin stability system
Accession: VEA63282
Location: 1017646-1017879
NCBI BlastP on this gene
Accession: VEA63282
Location: 1017646-1017879
NCBI BlastP on this gene
NCTC8900_01021
mak
Transcriptional regulatory protein OmpR
Accession: VEA63278
Location: 1015704-1016444
NCBI BlastP on this gene
Accession: VEA63278
Location: 1015704-1016444
NCBI BlastP on this gene
ompR_1
rbsK_1
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP013913 : Serratia fonticola strain GS2 Total score: 1.0 Cumulative Blast bit score: 266
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-dependent dsDNA exonuclease
Accession: ALX93791
Location: 2111520-2114771
NCBI BlastP on this gene
Accession: ALX93791
Location: 2111520-2114771
NCBI BlastP on this gene
AV650_09600
transcription antiterminator BglG
Accession: ALX93790
Location: 2110456-2111313
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 266
Sequence coverage: 100 %
E-value: 1e-84
NCBI BlastP on this gene
Accession: ALX93790
Location: 2110456-2111313
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 266
Sequence coverage: 100 %
E-value: 1e-84
NCBI BlastP on this gene
AV650_09595
AV650_09590
recombination associated protein
Accession: ALX93788
Location: 2108346-2109257
NCBI BlastP on this gene
Accession: ALX93788
Location: 2108346-2109257
NCBI BlastP on this gene
AV650_09585
AV650_09580
AV650_09575
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP014136 : Gibbsiella quercinecans strain FRB97 Total score: 1.0 Cumulative Blast bit score: 265
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AraC family transcriptional regulator
Accession: ATA19042
Location: 1468080-1469144
NCBI BlastP on this gene
Accession: ATA19042
Location: 1468080-1469144
NCBI BlastP on this gene
AWC35_06605
AWC35_06610
AWC35_06615
transcription antiterminator BglG
Accession: ATA19045
Location: 1472392-1473249
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 265
Sequence coverage: 101 %
E-value: 8e-84
NCBI BlastP on this gene
Accession: ATA19045
Location: 1472392-1473249
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 265
Sequence coverage: 101 %
E-value: 8e-84
NCBI BlastP on this gene
AWC35_06620
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP015613 : Serratia plymuthica PRI-2C chromosome Total score: 1.0 Cumulative Blast bit score: 264
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
sbcC
putative amino acid permease YhdG
Accession: ANS41430
Location: 1014894-1016261
NCBI BlastP on this gene
Accession: ANS41430
Location: 1014894-1016261
NCBI BlastP on this gene
yhdG
Transcription antiterminator LicT
Accession: ANS41429
Location: 1013777-1014619
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 264
Sequence coverage: 100 %
E-value: 2e-83
NCBI BlastP on this gene
Accession: ANS41429
Location: 1013777-1014619
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 264
Sequence coverage: 100 %
E-value: 2e-83
NCBI BlastP on this gene
licT_1
tRNA3(Ser)-specific nuclease WapA
Accession: ANS41428
Location: 1008588-1013459
NCBI BlastP on this gene
Accession: ANS41428
Location: 1008588-1013459
NCBI BlastP on this gene
wapA_1
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP000813 : Bacillus pumilus SAFR-032 chromosome Total score: 1.0 Cumulative Blast bit score: 262
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
transcription antiterminator LicT
Accession: ABV64211
Location: 3531245-3532078
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 262
Sequence coverage: 98 %
E-value: 7e-83
NCBI BlastP on this gene
Accession: ABV64211
Location: 3531245-3532078
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 262
Sequence coverage: 98 %
E-value: 7e-83
NCBI BlastP on this gene
BPUM_3567
PTS beta-glucoside transporter subunit IIA
Accession: ABV64210
Location: 3529010-3530917
NCBI BlastP on this gene
Accession: ABV64210
Location: 3529010-3530917
NCBI BlastP on this gene
BPUM_3566
BPUM_3565
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP031774 : Bacillus altitudinis strain Cr2-1 chromosome. Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ATP-binding cassette domain-containing protein
Accession: QDZ95456
Location: 2123073-2124671
NCBI BlastP on this gene
Accession: QDZ95456
Location: 2123073-2124671
NCBI BlastP on this gene
D0438_11080
energy-coupling factor transporter transmembrane protein EcfT
Accession: QDZ95455
Location: 2122349-2123083
NCBI BlastP on this gene
Accession: QDZ95455
Location: 2122349-2123083
NCBI BlastP on this gene
D0438_11075
D0438_11070
DUF1963 domain-containing protein
Accession: QDZ95453
Location: 2120371-2121186
NCBI BlastP on this gene
Accession: QDZ95453
Location: 2120371-2121186
NCBI BlastP on this gene
D0438_11065
PRD domain-containing protein
Accession: QDZ95452
Location: 2119313-2120146
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: QDZ95452
Location: 2119313-2120146
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
D0438_11060
PTS beta-glucoside transporter subunit EIIBCA
Accession: QDZ95451
Location: 2117080-2118987
NCBI BlastP on this gene
Accession: QDZ95451
Location: 2117080-2118987
NCBI BlastP on this gene
D0438_11055
glycoside hydrolase family 1 protein
Accession: QDZ95450
Location: 2115690-2117108
NCBI BlastP on this gene
Accession: QDZ95450
Location: 2115690-2117108
NCBI BlastP on this gene
D0438_11050
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP026008 : Bacillus aerophilus strain 232 chromosome Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
BAE_06675
energy-coupling factor transporter transmembrane protein EcfT
Accession: QAR52518
Location: 1267940-1268674
NCBI BlastP on this gene
Accession: QAR52518
Location: 1267940-1268674
NCBI BlastP on this gene
BAE_06670
BAE_06665
DUF1963 domain-containing protein
Accession: QAR52516
Location: 1265961-1266776
NCBI BlastP on this gene
Accession: QAR52516
Location: 1265961-1266776
NCBI BlastP on this gene
BAE_06660
PRD domain-containing protein
Accession: QAR52515
Location: 1264903-1265736
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: QAR52515
Location: 1264903-1265736
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
BAE_06655
PTS beta-glucoside transporter subunit EIIBCA
Accession: QAR52514
Location: 1262670-1264577
NCBI BlastP on this gene
Accession: QAR52514
Location: 1262670-1264577
NCBI BlastP on this gene
BAE_06650
glycoside hydrolase family 1 protein
Accession: QAR52513
Location: 1261280-1262698
NCBI BlastP on this gene
Accession: QAR52513
Location: 1261280-1262698
NCBI BlastP on this gene
BAE_06645
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP024204 : Bacillus altitudinis strain P-10 chromosome Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
transcription antiterminator LicT
Accession: ATP95885
Location: 3586300-3587133
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ATP95885
Location: 3586300-3587133
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
CSE15_18830
PTS beta-glucoside transporter subunit EIIBCA
Accession: ATP95884
Location: 3584067-3585974
NCBI BlastP on this gene
Accession: ATP95884
Location: 3584067-3585974
NCBI BlastP on this gene
CSE15_18825
glycoside hydrolase family 1 protein
Accession: ATP95883
Location: 3582677-3584095
NCBI BlastP on this gene
Accession: ATP95883
Location: 3582677-3584095
NCBI BlastP on this gene
CSE15_18820
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP018574 : Bacillus cellulasensis strain GLB197 Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
BS467_05290
BS467_05295
BS467_05300
BS467_05305
transcription antiterminator LicT
Accession: APP15176
Location: 1051308-1052141
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 1e-82
NCBI BlastP on this gene
Accession: APP15176
Location: 1051308-1052141
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 1e-82
NCBI BlastP on this gene
BS467_05310
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP012482 : Bacillus cellulasensis strain NJ-V2 Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AMR71_01320
AMR71_01325
AMR71_01330
AMR71_01335
transcription antiterminator LicT
Accession: ALM43939
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ALM43939
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
AMR71_01340
PTS beta-glucoside transporter subunit IIA
Accession: ALM43940
Location: 269336-271243
NCBI BlastP on this gene
Accession: ALM43940
Location: 269336-271243
NCBI BlastP on this gene
AMR71_01345
AMR71_01350
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP012330 : Bacillus cellulasensis strain NJ-V Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AKO66_01320
AKO66_01325
AKO66_01330
AKO66_01335
transcription antiterminator LicT
Accession: ANY95411
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ANY95411
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
AKO66_01340
PTS beta-glucoside transporter subunit IIA
Accession: ANY95412
Location: 269336-271243
NCBI BlastP on this gene
Accession: ANY95412
Location: 269336-271243
NCBI BlastP on this gene
AKO66_01345
AKO66_01350
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP012329 : Bacillus cellulasensis strain NJ-M2 Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
AKO65_05005
AKO65_05010
AKO65_05015
AKO65_05020
transcription antiterminator LicT
Accession: ALM27397
Location: 1024964-1025797
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ALM27397
Location: 1024964-1025797
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
AKO65_05025
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP011109 : Bacillus pumilus strain C4 genome. Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
transcription antiterminator LicT
Accession: ANT58472
Location: 3395899-3396732
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ANT58472
Location: 3395899-3396732
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
VP59_17195
PTS beta-glucoside transporter subunit IIA
Accession: ANT58471
Location: 3393666-3395573
NCBI BlastP on this gene
Accession: ANT58471
Location: 3393666-3395573
NCBI BlastP on this gene
VP59_17190
VP59_17185
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
CP009108 : Bacillus altitudinis strain GR-8 Total score: 1.0 Cumulative Blast bit score: 261
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
ID12_04170
ID12_04165
ID12_04160
ID12_04155
transcription antiterminator LicT
Accession: AKU30682
Location: 793084-793917
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: AKU30682
Location: 793084-793917
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
ID12_04150
PTS beta-glucoside transporter subunit IIA
Accession: AKU30681
Location: 790851-792758
NCBI BlastP on this gene
Accession: AKU30681
Location: 790851-792758
NCBI BlastP on this gene
ID12_04145
ID12_04140
Query: Pectobacterium carotovorum subsp. carotovorum BglY (bglY) and BglK
201. : AP021873 Serratia marcescens ATCC 274 DNA Total score: 1.0 Cumulative Blast bit score: 275
GH1
Location: 356-1780
Location: 356-1780
bglY
BglK
Location: 1874-2710
Location: 1874-2710
bglK
SMATCC274_08870
beta-glucoside operon transcriptional antiterminator
Accession: BBO61623
Location: 984528-985376
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 1e-87
NCBI BlastP on this gene
Accession: BBO61623
Location: 984528-985376
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 1e-87
NCBI BlastP on this gene
SMATCC274_08860
twitching motility protein PilT
Accession: BBO61622
Location: 984145-984531
NCBI BlastP on this gene
Accession: BBO61622
Location: 984145-984531
NCBI BlastP on this gene
SMATCC274_08850
SMATCC274_08840
SMATCC274_08830
recombination-associated protein RdgC
Accession: BBO61619
Location: 981769-982680
NCBI BlastP on this gene
Accession: BBO61619
Location: 981769-982680
NCBI BlastP on this gene
rdgC
SMATCC274_08810
202. : AP019009 Serratia marcescens AS-1 DNA Total score: 1.0 Cumulative Blast bit score: 275
sbcC
beta-glucoside operon transcriptional antiterminator
Accession: BBG68099
Location: 1018724-1019572
BlastP hit with bglK
Percentage identity: 49 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 4e-88
NCBI BlastP on this gene
Accession: BBG68099
Location: 1018724-1019572
BlastP hit with bglK
Percentage identity: 49 %
BlastP bit score: 275
Sequence coverage: 98 %
E-value: 4e-88
NCBI BlastP on this gene
SERAS_09020
twitching motility protein PilT
Accession: BBG68098
Location: 1018341-1018727
NCBI BlastP on this gene
Accession: BBG68098
Location: 1018341-1018727
NCBI BlastP on this gene
SERAS_09010
SERAS_09000
SERAS_08990
recombination-associated protein RdgC
Accession: BBG68095
Location: 1015965-1016876
NCBI BlastP on this gene
Accession: BBG68095
Location: 1015965-1016876
NCBI BlastP on this gene
rdgC
SERAS_08970
203. : CP047605 Serratia sp. NGAS9 chromosome Total score: 1.0 Cumulative Blast bit score: 274
sbcC
PRD domain-containing protein
Accession: QHI76891
Location: 977103-977951
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: QHI76891
Location: 977103-977951
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
GUC32_04605
GUC32_04600
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QHI76889
Location: 976490-976723
NCBI BlastP on this gene
Accession: QHI76889
Location: 976490-976723
NCBI BlastP on this gene
GUC32_04595
GUC32_04590
recombination-associated protein RdgC
Accession: QHI76887
Location: 974345-975256
NCBI BlastP on this gene
Accession: QHI76887
Location: 974345-975256
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QHI76886
Location: 973989-974279
NCBI BlastP on this gene
Accession: QHI76886
Location: 973989-974279
NCBI BlastP on this gene
ppnP
204. : CP041124 Serratia marcescens strain WVU-003 chromosome Total score: 1.0 Cumulative Blast bit score: 274
sbcC
PRD domain-containing protein
Accession: QDI22246
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: QDI22246
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
FBF90_04720
type II toxin-antitoxin system VapC family toxin
Accession: QDI22245
Location: 1011968-1012354
NCBI BlastP on this gene
Accession: QDI22245
Location: 1011968-1012354
NCBI BlastP on this gene
FBF90_04715
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QDI22244
Location: 1011738-1011971
NCBI BlastP on this gene
Accession: QDI22244
Location: 1011738-1011971
NCBI BlastP on this gene
FBF90_04710
FBF90_04705
recombination-associated protein RdgC
Accession: QDI22242
Location: 1009593-1010504
NCBI BlastP on this gene
Accession: QDI22242
Location: 1009593-1010504
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QDI22241
Location: 1009237-1009527
NCBI BlastP on this gene
Accession: QDI22241
Location: 1009237-1009527
NCBI BlastP on this gene
ppnP
205. : CP041122 Serratia marcescens strain WVU-001 chromosome Total score: 1.0 Cumulative Blast bit score: 274
sbcC
PRD domain-containing protein
Accession: QDI12503
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: QDI12503
Location: 1012351-1013199
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
FBF84_04720
type II toxin-antitoxin system VapC family toxin
Accession: QDI12502
Location: 1011968-1012354
NCBI BlastP on this gene
Accession: QDI12502
Location: 1011968-1012354
NCBI BlastP on this gene
FBF84_04715
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QDI12501
Location: 1011738-1011971
NCBI BlastP on this gene
Accession: QDI12501
Location: 1011738-1011971
NCBI BlastP on this gene
FBF84_04710
FBF84_04705
recombination-associated protein RdgC
Accession: QDI12499
Location: 1009593-1010504
NCBI BlastP on this gene
Accession: QDI12499
Location: 1009593-1010504
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QDI12498
Location: 1009237-1009527
NCBI BlastP on this gene
Accession: QDI12498
Location: 1009237-1009527
NCBI BlastP on this gene
ppnP
206. : CP018930 Serratia marcescens strain UMH12 chromosome Total score: 1.0 Cumulative Blast bit score: 274
BVG84_01565
transcription antiterminator BglG
Accession: ASM29766
Location: 322310-323158
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: ASM29766
Location: 322310-323158
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
BVG84_01560
twitching motility protein PilT
Accession: ASM29765
Location: 321927-322313
NCBI BlastP on this gene
Accession: ASM29765
Location: 321927-322313
NCBI BlastP on this gene
BVG84_01555
BVG84_01550
BVG84_01545
recombination-associated protein RdgC
Accession: ASM29762
Location: 319552-320463
NCBI BlastP on this gene
Accession: ASM29762
Location: 319552-320463
NCBI BlastP on this gene
BVG84_01540
BVG84_01535
207. : CP018915 Serratia marcescens strain UMH1 chromosome Total score: 1.0 Cumulative Blast bit score: 274
BVG95_01540
transcription antiterminator BglG
Accession: ASL81665
Location: 319631-320479
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
Accession: ASL81665
Location: 319631-320479
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 274
Sequence coverage: 98 %
E-value: 2e-87
NCBI BlastP on this gene
BVG95_01535
twitching motility protein PilT
Accession: ASL81664
Location: 319248-319634
NCBI BlastP on this gene
Accession: ASL81664
Location: 319248-319634
NCBI BlastP on this gene
BVG95_01530
BVG95_01525
BVG95_01520
recombination-associated protein RdgC
Accession: ASL81661
Location: 316873-317784
NCBI BlastP on this gene
Accession: ASL81661
Location: 316873-317784
NCBI BlastP on this gene
BVG95_01515
BVG95_01510
208. : LT906479 Serratia ficaria strain NCTC12148 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 273
serine/threonine protein kinase
Accession: SNV90830
Location: 1140607-1141617
NCBI BlastP on this gene
Accession: SNV90830
Location: 1140607-1141617
NCBI BlastP on this gene
SAMEA4384070_01073
4-aminobutyrate aminotransferase GabT
Accession: SNV90820
Location: 1139047-1140354
NCBI BlastP on this gene
Accession: SNV90820
Location: 1139047-1140354
NCBI BlastP on this gene
gabT
Low-affinity putrescine importer PlaP
Accession: SNV90812
Location: 1137617-1138984
NCBI BlastP on this gene
Accession: SNV90812
Location: 1137617-1138984
NCBI BlastP on this gene
plaP_1
Transcription antiterminator LicT
Accession: SNV90802
Location: 1136588-1137430
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 273
Sequence coverage: 100 %
E-value: 3e-87
NCBI BlastP on this gene
Accession: SNV90802
Location: 1136588-1137430
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 273
Sequence coverage: 100 %
E-value: 3e-87
NCBI BlastP on this gene
licT_1
mak
Transcriptional regulatory protein OmpR
Accession: SNV90786
Location: 1134646-1135380
NCBI BlastP on this gene
Accession: SNV90786
Location: 1134646-1135380
NCBI BlastP on this gene
ompR_1
ABC-type sugar transport system, auxiliary component
Accession: SNV90777
Location: 1133904-1134590
NCBI BlastP on this gene
Accession: SNV90777
Location: 1133904-1134590
NCBI BlastP on this gene
SAMEA4384070_01067
D-ribose-binding periplasmic protein precursor
Accession: SNV90768
Location: 1132904-1133845
NCBI BlastP on this gene
Accession: SNV90768
Location: 1132904-1133845
NCBI BlastP on this gene
rbsB_2
209. : CP047391 Serratia marcescens strain 1602 chromosome Total score: 1.0 Cumulative Blast bit score: 272
sbcC
PRD domain-containing protein
Accession: QHE55762
Location: 983893-984741
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: QHE55762
Location: 983893-984741
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
GRX59_04605
GRX59_04600
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: QHE55760
Location: 983280-983513
NCBI BlastP on this gene
Accession: QHE55760
Location: 983280-983513
NCBI BlastP on this gene
GRX59_04595
GRX59_04590
recombination-associated protein RdgC
Accession: QHE55758
Location: 981135-982046
NCBI BlastP on this gene
Accession: QHE55758
Location: 981135-982046
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QHE55757
Location: 980779-981069
NCBI BlastP on this gene
Accession: QHE55757
Location: 980779-981069
NCBI BlastP on this gene
ppnP
210. : CP029746 Serratia marcescens strain AR_0122 Total score: 1.0 Cumulative Blast bit score: 272
AM369_17260
PRD domain-containing protein
Accession: AWS59912
Location: 3659040-3659888
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWS59912
Location: 3659040-3659888
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM369_17255
AM369_17250
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWS59910
Location: 3658427-3658660
NCBI BlastP on this gene
Accession: AWS59910
Location: 3658427-3658660
NCBI BlastP on this gene
AM369_17245
AM369_17240
recombination-associated protein RdgC
Accession: AWS59908
Location: 3656282-3657193
NCBI BlastP on this gene
Accession: AWS59908
Location: 3656282-3657193
NCBI BlastP on this gene
AM369_17235
pyrimidine/purine nucleoside phosphorylase
Accession: AWS59907
Location: 3655926-3656216
NCBI BlastP on this gene
Accession: AWS59907
Location: 3655926-3656216
NCBI BlastP on this gene
AM369_17230
211. : CP029715 Serratia marcescens strain AR_0131 chromosome Total score: 1.0 Cumulative Blast bit score: 272
PRD domain-containing protein
Accession: AWS71293
Location: 5081706-5082554
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWS71293
Location: 5081706-5082554
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM378_24130
AM378_24125
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWS71291
Location: 5081093-5081326
NCBI BlastP on this gene
Accession: AWS71291
Location: 5081093-5081326
NCBI BlastP on this gene
AM378_24120
AM378_24115
recombination-associated protein RdgC
Accession: AWS71289
Location: 5078948-5079859
NCBI BlastP on this gene
Accession: AWS71289
Location: 5078948-5079859
NCBI BlastP on this gene
AM378_24110
pyrimidine/purine nucleoside phosphorylase
Accession: AWS71288
Location: 5078592-5078882
NCBI BlastP on this gene
Accession: AWS71288
Location: 5078592-5078882
NCBI BlastP on this gene
AM378_24105
212. : CP028949 Serratia marcescens strain AR_0121 chromosome Total score: 1.0 Cumulative Blast bit score: 272
AM368_06645
PRD domain-containing protein
Accession: AWC69937
Location: 1377712-1378560
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWC69937
Location: 1377712-1378560
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM368_06650
AM368_06655
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWC69939
Location: 1378940-1379173
NCBI BlastP on this gene
Accession: AWC69939
Location: 1378940-1379173
NCBI BlastP on this gene
AM368_06660
AM368_06665
recombination-associated protein RdgC
Accession: AWC69941
Location: 1380407-1381318
NCBI BlastP on this gene
Accession: AWC69941
Location: 1380407-1381318
NCBI BlastP on this gene
AM368_06670
pyrimidine/purine nucleoside phosphorylase
Accession: AWC69942
Location: 1381384-1381674
NCBI BlastP on this gene
Accession: AWC69942
Location: 1381384-1381674
NCBI BlastP on this gene
AM368_06675
213. : CP028948 Serratia marcescens strain AR_0123 chromosome Total score: 1.0 Cumulative Blast bit score: 272
AM370_02385
PRD domain-containing protein
Accession: AWC87888
Location: 477583-478431
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AWC87888
Location: 477583-478431
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 272
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AM370_02390
AM370_02395
type II toxin-antitoxin system prevent-host-death family antitoxin
Accession: AWC87890
Location: 478811-479044
NCBI BlastP on this gene
Accession: AWC87890
Location: 478811-479044
NCBI BlastP on this gene
AM370_02400
AM370_02405
recombination-associated protein RdgC
Accession: AWC87892
Location: 480278-481189
NCBI BlastP on this gene
Accession: AWC87892
Location: 480278-481189
NCBI BlastP on this gene
AM370_02410
pyrimidine/purine nucleoside phosphorylase
Accession: AWC87893
Location: 481255-481545
NCBI BlastP on this gene
Accession: AWC87893
Location: 481255-481545
NCBI BlastP on this gene
AM370_02415
214. : LT883155 Serratia grimesii isolate BXF1 genome assembly, chromosome. Total score: 1.0 Cumulative Blast bit score: 271
sbcC
Transcription antiterminator LicT
Accession: SMZ55352
Location: 1038009-1038851
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: SMZ55352
Location: 1038009-1038851
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 3e-86
NCBI BlastP on this gene
licT_1
mak
Transcriptional regulatory protein OmpR
Accession: SMZ55350
Location: 1036071-1036811
NCBI BlastP on this gene
Accession: SMZ55350
Location: 1036071-1036811
NCBI BlastP on this gene
ompR_1
ydjH_1
putative fructose-bisphosphate aldolase
Accession: SMZ55348
Location: 1034223-1035083
NCBI BlastP on this gene
Accession: SMZ55348
Location: 1034223-1035083
NCBI BlastP on this gene
fbaA_1
215. : CP050171 Serratia fonticola strain CPSE11 chromosome Total score: 1.0 Cumulative Blast bit score: 271
HAP32_03519
transcription antiterminator BglG
Accession: QIP92998
Location: 3898814-3899671
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: QIP92998
Location: 3898814-3899671
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
HAP32_03518
HAP32_03517
DNA recombination-dependent growth factor C
Accession: QIP92996
Location: 3896565-3897476
NCBI BlastP on this gene
Accession: QIP92996
Location: 3896565-3897476
NCBI BlastP on this gene
HAP32_03516
HAP32_03515
HAP32_03514
216. : CP040182 Serratia fonticola strain MS5 chromosome Total score: 1.0 Cumulative Blast bit score: 271
sbcC
PRD domain-containing protein
Accession: QCR60083
Location: 1508365-1509222
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: QCR60083
Location: 1508365-1509222
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
FD644_06815
FD644_06810
recombination-associated protein RdgC
Accession: QCR60081
Location: 1506126-1507037
NCBI BlastP on this gene
Accession: QCR60081
Location: 1506126-1507037
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: QCR60080
Location: 1505759-1506049
NCBI BlastP on this gene
Accession: QCR60080
Location: 1505759-1506049
NCBI BlastP on this gene
ppnP
DUF1177 domain-containing protein
Accession: QCR60079
Location: 1504645-1505574
NCBI BlastP on this gene
Accession: QCR60079
Location: 1504645-1505574
NCBI BlastP on this gene
FD644_06795
217. : CP031316 Serratia marcescens strain N4-5 chromosome Total score: 1.0 Cumulative Blast bit score: 271
Exonuclease SbcC family protein
Accession: AXK22751
Location: 993288-996539
NCBI BlastP on this gene
Accession: AXK22751
Location: 993288-996539
NCBI BlastP on this gene
SmN45_0945
Transcription antiterminator LicT
Accession: AXK22750
Location: 992208-993056
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AXK22750
Location: 992208-993056
BlastP hit with bglK
Percentage identity: 48 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 1e-86
NCBI BlastP on this gene
licT
SmN45_0943
Prevent-host-death family protein
Accession: AXK22748
Location: 991595-991828
NCBI BlastP on this gene
Accession: AXK22748
Location: 991595-991828
NCBI BlastP on this gene
SmN45_0942
SmN45_0941
Recombination-associated protein RdgC
Accession: AXK22746
Location: 989449-990360
NCBI BlastP on this gene
Accession: AXK22746
Location: 989449-990360
NCBI BlastP on this gene
rdgC
SmN45_0939
218. : CP023956 Serratia fonticola strain FDAARGOS_411 chromosome Total score: 1.0 Cumulative Blast bit score: 271
CRN79_04260
transcription antiterminator BglG
Accession: ATM75107
Location: 960943-961800
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 2e-86
NCBI BlastP on this gene
Accession: ATM75107
Location: 960943-961800
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 2e-86
NCBI BlastP on this gene
CRN79_04265
CRN79_04270
recombination-associated protein RdgC
Accession: ATM75109
Location: 963009-963920
NCBI BlastP on this gene
Accession: ATM75109
Location: 963009-963920
NCBI BlastP on this gene
CRN79_04275
pyrimidine/purine nucleoside phosphorylase
Accession: ATM75110
Location: 963997-964287
NCBI BlastP on this gene
Accession: ATM75110
Location: 963997-964287
NCBI BlastP on this gene
CRN79_04280
CRN79_04285
219. : CP014017 Serratia liquefaciens strain FDAARGOS_125 chromosome Total score: 1.0 Cumulative Blast bit score: 271
AL485_07500
PRD domain-containing protein
Accession: AMG99026
Location: 1582508-1583350
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
Accession: AMG99026
Location: 1582508-1583350
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 100 %
E-value: 1e-86
NCBI BlastP on this gene
AL485_07505
220. : CP011254 Serratia fonticola strain DSM 4576 Total score: 1.0 Cumulative Blast bit score: 271
ATP-dependent dsDNA exonuclease
Accession: AKG70130
Location: 3169195-3172446
NCBI BlastP on this gene
Accession: AKG70130
Location: 3169195-3172446
NCBI BlastP on this gene
WN53_14050
transcription antiterminator BglG
Accession: AKG70129
Location: 3168131-3168988
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
Accession: AKG70129
Location: 3168131-3168988
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 3e-86
NCBI BlastP on this gene
WN53_14045
WN53_14040
recombination associated protein
Accession: AKG70127
Location: 3166021-3166932
NCBI BlastP on this gene
Accession: AKG70127
Location: 3166021-3166932
NCBI BlastP on this gene
WN53_14035
WN53_14030
WN53_14025
221. : CP006252 Serratia liquefaciens ATCC 27592 Total score: 1.0 Cumulative Blast bit score: 271
ATP-dependent dsDNA exonuclease
Accession: AGQ29717
Location: 959249-962500
NCBI BlastP on this gene
Accession: AGQ29717
Location: 959249-962500
NCBI BlastP on this gene
M495_04455
beta-1,4-xylanase
Accession: AGQ29716
Location: 958211-959077
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: AGQ29716
Location: 958211-959077
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 271
Sequence coverage: 98 %
E-value: 4e-86
NCBI BlastP on this gene
M495_04450
M495_04445
M495_04440
M495_04435
M495_04430
M495_04425
222. : LS483469 Serratia plymuthica strain NCTC12961 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 270
sbcC_2
sbcC_1
HTH-type transcriptional regulator gadW
Accession: SQI32688
Location: 1059059-1059520
NCBI BlastP on this gene
Accession: SQI32688
Location: 1059059-1059520
NCBI BlastP on this gene
gadW
Proline-specific permease ProY
Accession: SQI32685
Location: 1057421-1058788
NCBI BlastP on this gene
Accession: SQI32685
Location: 1057421-1058788
NCBI BlastP on this gene
proY_2
Transcription antiterminator LicT
Accession: SQI32683
Location: 1056304-1057146
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: SQI32683
Location: 1056304-1057146
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
licT_1
NCTC12961_01230
Antitoxin of toxin-antitoxin stability system
Accession: SQI32681
Location: 1055685-1055918
NCBI BlastP on this gene
Accession: SQI32681
Location: 1055685-1055918
NCBI BlastP on this gene
NCTC12961_01229
mak
Transcriptional regulatory protein OmpR
Accession: SQI32679
Location: 1053742-1054482
NCBI BlastP on this gene
Accession: SQI32679
Location: 1053742-1054482
NCBI BlastP on this gene
ompR_1
NCTC12961_01226
223. : CP033893 Serratia liquefaciens strain FG3 chromosome Total score: 1.0 Cumulative Blast bit score: 270
sbcC
PRD domain-containing protein
Accession: QDL35495
Location: 3682954-3683796
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: QDL35495
Location: 3682954-3683796
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
EGO53_18000
224. : CP033162 Serratia sp. P2ACOL2 chromosome Total score: 1.0 Cumulative Blast bit score: 270
sbcC
PRD domain-containing protein
Accession: AYO40686
Location: 831433-832275
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
Accession: AYO40686
Location: 831433-832275
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 4e-86
NCBI BlastP on this gene
EBA31_04005
EBA31_04000
EBA31_03995
EBA31_03990
carbohydrate kinase family protein
Accession: AYO36504
Location: 828328-829341
NCBI BlastP on this gene
Accession: AYO36504
Location: 828328-829341
NCBI BlastP on this gene
EBA31_03985
ketose 1,6-bisphosphate aldolase
Accession: AYO36503
Location: 827468-828328
NCBI BlastP on this gene
Accession: AYO36503
Location: 827468-828328
NCBI BlastP on this gene
EBA31_03980
225. : CP033055 Serratia sp. 3ACOL1 chromosome Total score: 1.0 Cumulative Blast bit score: 270
PRD domain-containing protein
Accession: AYM93815
Location: 5832446-5833303
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 270
Sequence coverage: 98 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AYM93815
Location: 5832446-5833303
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 270
Sequence coverage: 98 %
E-value: 8e-86
NCBI BlastP on this gene
D9980_26435
D9980_26430
recombination-associated protein RdgC
Accession: AYM93813
Location: 5830336-5831247
NCBI BlastP on this gene
Accession: AYM93813
Location: 5830336-5831247
NCBI BlastP on this gene
rdgC
pyrimidine/purine nucleoside phosphorylase
Accession: AYM93812
Location: 5829969-5830259
NCBI BlastP on this gene
Accession: AYM93812
Location: 5829969-5830259
NCBI BlastP on this gene
D9980_26420
DUF1177 domain-containing protein
Accession: AYM93811
Location: 5828855-5829784
NCBI BlastP on this gene
Accession: AYM93811
Location: 5828855-5829784
NCBI BlastP on this gene
D9980_26415
226. : CP012097 Serratia plymuthica strain 3Re4-18 Total score: 1.0 Cumulative Blast bit score: 270
ATP-dependent dsDNA exonuclease
Accession: ANJ97339
Location: 1092870-1096124
NCBI BlastP on this gene
Accession: ANJ97339
Location: 1092870-1096124
NCBI BlastP on this gene
ADP73_05040
ADP73_05035
ADP73_05030
transcription antiterminator BglG
Accession: ANJ97337
Location: 1089659-1090501
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: ANJ97337
Location: 1089659-1090501
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
ADP73_05025
ADP73_05020
ADP73_05015
ADP73_05010
ADP73_05005
227. : CP012096 Serratia plymuthica strain 3Rp8 Total score: 1.0 Cumulative Blast bit score: 270
ATP-dependent dsDNA exonuclease
Accession: ANJ92596
Location: 1264435-1267689
NCBI BlastP on this gene
Accession: ANJ92596
Location: 1264435-1267689
NCBI BlastP on this gene
ADP72_06195
ADP72_06200
ADP72_06205
transcription antiterminator BglG
Accession: ANJ92598
Location: 1270058-1270900
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: ANJ92598
Location: 1270058-1270900
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
ADP72_06210
ADP72_06215
ADP72_06220
ADP72_06225
ADP72_06230
228. : CP007439 Serratia plymuthica strain V4 genome. Total score: 1.0 Cumulative Blast bit score: 270
ATP-dependent dsDNA exonuclease
Accession: AHY05951
Location: 1071599-1074853
NCBI BlastP on this gene
Accession: AHY05951
Location: 1071599-1074853
NCBI BlastP on this gene
sch_05035
sch_05030
sch_05025
transcription antiterminator BglG
Accession: AHY05948
Location: 1068388-1069230
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AHY05948
Location: 1068388-1069230
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
sch_05020
sch_05015
sch_05010
sch_05005
sch_05000
229. : CP006566 Serratia plymuthica S13 Total score: 1.0 Cumulative Blast bit score: 270
ATP-dependent dsDNA exonuclease
Accession: AGP43241
Location: 1039188-1042442
NCBI BlastP on this gene
Accession: AGP43241
Location: 1039188-1042442
NCBI BlastP on this gene
M621_04885
M621_04880
M621_04875
beta-1,4-xylanase
Accession: AGP43239
Location: 1035977-1036819
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AGP43239
Location: 1035977-1036819
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
M621_04870
M621_04865
M621_04860
M621_04855
M621_04850
230. : CP006250 Serratia plymuthica 4Rx13 Total score: 1.0 Cumulative Blast bit score: 270
sbcC
transcriptional regulatory protein
Accession: AGO53887
Location: 972639-973100
NCBI BlastP on this gene
Accession: AGO53887
Location: 972639-973100
NCBI BlastP on this gene
SOD_c08900
SOD_c08890
beta-glucoside operon antiterminator
Accession: AGO53885
Location: 969887-970729
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
Accession: AGO53885
Location: 969887-970729
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 270
Sequence coverage: 100 %
E-value: 8e-86
NCBI BlastP on this gene
arbG
mak
transcriptional regulatory protein OmpR
Accession: AGO53883
Location: 967947-968687
NCBI BlastP on this gene
Accession: AGO53883
Location: 967947-968687
NCBI BlastP on this gene
ompR1
ribokinase-like domain-containing protein
Accession: AGO53882
Location: 966959-967972
NCBI BlastP on this gene
Accession: AGO53882
Location: 966959-967972
NCBI BlastP on this gene
SOD_c08850
D-tagatose-1,6-bisphosphate aldolase subunit GatY
Accession: AGO53881
Location: 966099-966959
NCBI BlastP on this gene
Accession: AGO53881
Location: 966099-966959
NCBI BlastP on this gene
gatY1
231. : CP048784 Serratia liquefaciens strain S1 chromosome Total score: 1.0 Cumulative Blast bit score: 269
sbcC
PRD domain-containing protein
Accession: QIC85739
Location: 997649-998491
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
Accession: QIC85739
Location: 997649-998491
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
F0336_04650
type II toxin-antitoxin system VapC family toxin
Accession: F0336_04645
Location: 997420-997562
NCBI BlastP on this gene
Accession: F0336_04645
Location: 997420-997562
NCBI BlastP on this gene
F0336_04645
mak
response regulator transcription factor
Accession: QIC85737
Location: 995466-996206
NCBI BlastP on this gene
Accession: QIC85737
Location: 995466-996206
NCBI BlastP on this gene
F0336_04635
carbohydrate kinase family protein
Accession: QIC85736
Location: 994478-995491
NCBI BlastP on this gene
Accession: QIC85736
Location: 994478-995491
NCBI BlastP on this gene
F0336_04630
ketose 1,6-bisphosphate aldolase
Accession: QIC85735
Location: 993618-994478
NCBI BlastP on this gene
Accession: QIC85735
Location: 993618-994478
NCBI BlastP on this gene
F0336_04625
232. : CP011303 Serratia liquefaciens strain HUMV-21 Total score: 1.0 Cumulative Blast bit score: 269
ATP-dependent dsDNA exonuclease
Accession: AKE11835
Location: 4016221-4019472
NCBI BlastP on this gene
Accession: AKE11835
Location: 4016221-4019472
NCBI BlastP on this gene
XJ20_18795
transcription antiterminator BglG
Accession: AKE13125
Location: 4019668-4020510
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
Accession: AKE13125
Location: 4019668-4020510
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 2e-85
NCBI BlastP on this gene
XJ20_18800
233. : CP002775 Serratia sp. AS13 Total score: 1.0 Cumulative Blast bit score: 269
SerAS13_0964
SerAS13_0962
transcriptional antiterminator, BglG
Accession: AEG26773
Location: 1060650-1061492
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
Accession: AEG26773
Location: 1060650-1061492
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
SerAS13_0961
SerAS13_0960
prevent-host-death family protein
Accession: AEG26771
Location: 1060031-1060264
NCBI BlastP on this gene
Accession: AEG26771
Location: 1060031-1060264
NCBI BlastP on this gene
SerAS13_0959
SerAS13_0958
two component transcriptional regulator, winged helix family
Accession: AEG26769
Location: 1058089-1058829
NCBI BlastP on this gene
Accession: AEG26769
Location: 1058089-1058829
NCBI BlastP on this gene
SerAS13_0957
SerAS13_0956
234. : CP002774 Serratia sp. AS12 Total score: 1.0 Cumulative Blast bit score: 269
SerAS12_0964
SerAS12_0962
transcriptional antiterminator, BglG
Accession: AEF49065
Location: 1060653-1061495
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
Accession: AEF49065
Location: 1060653-1061495
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
SerAS12_0961
SerAS12_0960
prevent-host-death family protein
Accession: AEF49063
Location: 1060034-1060267
NCBI BlastP on this gene
Accession: AEF49063
Location: 1060034-1060267
NCBI BlastP on this gene
SerAS12_0959
SerAS12_0958
two component transcriptional regulator, winged helix family
Accession: AEF49061
Location: 1058092-1058832
NCBI BlastP on this gene
Accession: AEF49061
Location: 1058092-1058832
NCBI BlastP on this gene
SerAS12_0957
SerAS12_0956
235. : CP002773 Serratia plymuthica AS9 Total score: 1.0 Cumulative Blast bit score: 269
SerAS9_0964
SerAS9_0962
transcriptional antiterminator, BglG
Accession: AEF44113
Location: 1060838-1061680
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
Accession: AEF44113
Location: 1060838-1061680
BlastP hit with bglK
Percentage identity: 47 %
BlastP bit score: 269
Sequence coverage: 100 %
E-value: 1e-85
NCBI BlastP on this gene
SerAS9_0961
SerAS9_0960
prevent-host-death family protein
Accession: AEF44111
Location: 1060219-1060452
NCBI BlastP on this gene
Accession: AEF44111
Location: 1060219-1060452
NCBI BlastP on this gene
SerAS9_0959
SerAS9_0958
two component transcriptional regulator, winged helix family
Accession: AEF44109
Location: 1058277-1059017
NCBI BlastP on this gene
Accession: AEF44109
Location: 1058277-1059017
NCBI BlastP on this gene
SerAS9_0957
SerAS9_0956
236. : LR134478 Serratia plymuthica strain NCTC8015 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 267
sbcC
Low-affinity putrescine importer PlaP
Accession: VEI16987
Location: 1030649-1032016
NCBI BlastP on this gene
Accession: VEI16987
Location: 1030649-1032016
NCBI BlastP on this gene
plaP_1
Transcription antiterminator LicT
Accession: VEI16985
Location: 1029533-1030375
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
Accession: VEI16985
Location: 1029533-1030375
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
licT_1
NCTC8015_00990
Antitoxin of toxin-antitoxin stability system
Accession: VEI16981
Location: 1028914-1029147
NCBI BlastP on this gene
Accession: VEI16981
Location: 1028914-1029147
NCBI BlastP on this gene
NCTC8015_00989
mak
Transcriptional regulatory protein OmpR
Accession: VEI16977
Location: 1026971-1027711
NCBI BlastP on this gene
Accession: VEI16977
Location: 1026971-1027711
NCBI BlastP on this gene
ompR_1
rbsK_1
237. : LR134151 Serratia plymuthica strain NCTC8900 genome assembly, chromosome: 1. Total score: 1.0 Cumulative Blast bit score: 267
sbcC_1
Low-affinity putrescine importer PlaP
Accession: VEA63288
Location: 1019379-1020746
NCBI BlastP on this gene
Accession: VEA63288
Location: 1019379-1020746
NCBI BlastP on this gene
plaP_1
Transcription antiterminator LicT
Accession: VEA63286
Location: 1018263-1019105
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
Accession: VEA63286
Location: 1018263-1019105
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 267
Sequence coverage: 100 %
E-value: 6e-85
NCBI BlastP on this gene
licT_1
NCTC8900_01022
Antitoxin of toxin-antitoxin stability system
Accession: VEA63282
Location: 1017646-1017879
NCBI BlastP on this gene
Accession: VEA63282
Location: 1017646-1017879
NCBI BlastP on this gene
NCTC8900_01021
mak
Transcriptional regulatory protein OmpR
Accession: VEA63278
Location: 1015704-1016444
NCBI BlastP on this gene
Accession: VEA63278
Location: 1015704-1016444
NCBI BlastP on this gene
ompR_1
rbsK_1
238. : CP013913 Serratia fonticola strain GS2 Total score: 1.0 Cumulative Blast bit score: 266
ATP-dependent dsDNA exonuclease
Accession: ALX93791
Location: 2111520-2114771
NCBI BlastP on this gene
Accession: ALX93791
Location: 2111520-2114771
NCBI BlastP on this gene
AV650_09600
transcription antiterminator BglG
Accession: ALX93790
Location: 2110456-2111313
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 266
Sequence coverage: 100 %
E-value: 1e-84
NCBI BlastP on this gene
Accession: ALX93790
Location: 2110456-2111313
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 266
Sequence coverage: 100 %
E-value: 1e-84
NCBI BlastP on this gene
AV650_09595
AV650_09590
recombination associated protein
Accession: ALX93788
Location: 2108346-2109257
NCBI BlastP on this gene
Accession: ALX93788
Location: 2108346-2109257
NCBI BlastP on this gene
AV650_09585
AV650_09580
AV650_09575
239. : CP014136 Gibbsiella quercinecans strain FRB97 Total score: 1.0 Cumulative Blast bit score: 265
AraC family transcriptional regulator
Accession: ATA19042
Location: 1468080-1469144
NCBI BlastP on this gene
Accession: ATA19042
Location: 1468080-1469144
NCBI BlastP on this gene
AWC35_06605
AWC35_06610
AWC35_06615
transcription antiterminator BglG
Accession: ATA19045
Location: 1472392-1473249
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 265
Sequence coverage: 101 %
E-value: 8e-84
NCBI BlastP on this gene
Accession: ATA19045
Location: 1472392-1473249
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 265
Sequence coverage: 101 %
E-value: 8e-84
NCBI BlastP on this gene
AWC35_06620
240. : CP015613 Serratia plymuthica PRI-2C chromosome Total score: 1.0 Cumulative Blast bit score: 264
sbcC
putative amino acid permease YhdG
Accession: ANS41430
Location: 1014894-1016261
NCBI BlastP on this gene
Accession: ANS41430
Location: 1014894-1016261
NCBI BlastP on this gene
yhdG
Transcription antiterminator LicT
Accession: ANS41429
Location: 1013777-1014619
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 264
Sequence coverage: 100 %
E-value: 2e-83
NCBI BlastP on this gene
Accession: ANS41429
Location: 1013777-1014619
BlastP hit with bglK
Percentage identity: 46 %
BlastP bit score: 264
Sequence coverage: 100 %
E-value: 2e-83
NCBI BlastP on this gene
licT_1
tRNA3(Ser)-specific nuclease WapA
Accession: ANS41428
Location: 1008588-1013459
NCBI BlastP on this gene
Accession: ANS41428
Location: 1008588-1013459
NCBI BlastP on this gene
wapA_1
241. : CP000813 Bacillus pumilus SAFR-032 chromosome Total score: 1.0 Cumulative Blast bit score: 262
transcription antiterminator LicT
Accession: ABV64211
Location: 3531245-3532078
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 262
Sequence coverage: 98 %
E-value: 7e-83
NCBI BlastP on this gene
Accession: ABV64211
Location: 3531245-3532078
BlastP hit with bglK
Percentage identity: 45 %
BlastP bit score: 262
Sequence coverage: 98 %
E-value: 7e-83
NCBI BlastP on this gene
BPUM_3567
PTS beta-glucoside transporter subunit IIA
Accession: ABV64210
Location: 3529010-3530917
NCBI BlastP on this gene
Accession: ABV64210
Location: 3529010-3530917
NCBI BlastP on this gene
BPUM_3566
BPUM_3565
242. : CP031774 Bacillus altitudinis strain Cr2-1 chromosome. Total score: 1.0 Cumulative Blast bit score: 261
ATP-binding cassette domain-containing protein
Accession: QDZ95456
Location: 2123073-2124671
NCBI BlastP on this gene
Accession: QDZ95456
Location: 2123073-2124671
NCBI BlastP on this gene
D0438_11080
energy-coupling factor transporter transmembrane protein EcfT
Accession: QDZ95455
Location: 2122349-2123083
NCBI BlastP on this gene
Accession: QDZ95455
Location: 2122349-2123083
NCBI BlastP on this gene
D0438_11075
D0438_11070
DUF1963 domain-containing protein
Accession: QDZ95453
Location: 2120371-2121186
NCBI BlastP on this gene
Accession: QDZ95453
Location: 2120371-2121186
NCBI BlastP on this gene
D0438_11065
PRD domain-containing protein
Accession: QDZ95452
Location: 2119313-2120146
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: QDZ95452
Location: 2119313-2120146
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
D0438_11060
PTS beta-glucoside transporter subunit EIIBCA
Accession: QDZ95451
Location: 2117080-2118987
NCBI BlastP on this gene
Accession: QDZ95451
Location: 2117080-2118987
NCBI BlastP on this gene
D0438_11055
glycoside hydrolase family 1 protein
Accession: QDZ95450
Location: 2115690-2117108
NCBI BlastP on this gene
Accession: QDZ95450
Location: 2115690-2117108
NCBI BlastP on this gene
D0438_11050
243. : CP026008 Bacillus aerophilus strain 232 chromosome Total score: 1.0 Cumulative Blast bit score: 261
BAE_06675
energy-coupling factor transporter transmembrane protein EcfT
Accession: QAR52518
Location: 1267940-1268674
NCBI BlastP on this gene
Accession: QAR52518
Location: 1267940-1268674
NCBI BlastP on this gene
BAE_06670
BAE_06665
DUF1963 domain-containing protein
Accession: QAR52516
Location: 1265961-1266776
NCBI BlastP on this gene
Accession: QAR52516
Location: 1265961-1266776
NCBI BlastP on this gene
BAE_06660
PRD domain-containing protein
Accession: QAR52515
Location: 1264903-1265736
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: QAR52515
Location: 1264903-1265736
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
BAE_06655
PTS beta-glucoside transporter subunit EIIBCA
Accession: QAR52514
Location: 1262670-1264577
NCBI BlastP on this gene
Accession: QAR52514
Location: 1262670-1264577
NCBI BlastP on this gene
BAE_06650
glycoside hydrolase family 1 protein
Accession: QAR52513
Location: 1261280-1262698
NCBI BlastP on this gene
Accession: QAR52513
Location: 1261280-1262698
NCBI BlastP on this gene
BAE_06645
244. : CP024204 Bacillus altitudinis strain P-10 chromosome Total score: 1.0 Cumulative Blast bit score: 261
transcription antiterminator LicT
Accession: ATP95885
Location: 3586300-3587133
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ATP95885
Location: 3586300-3587133
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
CSE15_18830
PTS beta-glucoside transporter subunit EIIBCA
Accession: ATP95884
Location: 3584067-3585974
NCBI BlastP on this gene
Accession: ATP95884
Location: 3584067-3585974
NCBI BlastP on this gene
CSE15_18825
glycoside hydrolase family 1 protein
Accession: ATP95883
Location: 3582677-3584095
NCBI BlastP on this gene
Accession: ATP95883
Location: 3582677-3584095
NCBI BlastP on this gene
CSE15_18820
245. : CP018574 Bacillus cellulasensis strain GLB197 Total score: 1.0 Cumulative Blast bit score: 261
BS467_05290
BS467_05295
BS467_05300
BS467_05305
transcription antiterminator LicT
Accession: APP15176
Location: 1051308-1052141
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 1e-82
NCBI BlastP on this gene
Accession: APP15176
Location: 1051308-1052141
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 1e-82
NCBI BlastP on this gene
BS467_05310
246. : CP012482 Bacillus cellulasensis strain NJ-V2 Total score: 1.0 Cumulative Blast bit score: 261
AMR71_01320
AMR71_01325
AMR71_01330
AMR71_01335
transcription antiterminator LicT
Accession: ALM43939
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ALM43939
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
AMR71_01340
PTS beta-glucoside transporter subunit IIA
Accession: ALM43940
Location: 269336-271243
NCBI BlastP on this gene
Accession: ALM43940
Location: 269336-271243
NCBI BlastP on this gene
AMR71_01345
AMR71_01350
247. : CP012330 Bacillus cellulasensis strain NJ-V Total score: 1.0 Cumulative Blast bit score: 261
AKO66_01320
AKO66_01325
AKO66_01330
AKO66_01335
transcription antiterminator LicT
Accession: ANY95411
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ANY95411
Location: 268177-269010
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
AKO66_01340
PTS beta-glucoside transporter subunit IIA
Accession: ANY95412
Location: 269336-271243
NCBI BlastP on this gene
Accession: ANY95412
Location: 269336-271243
NCBI BlastP on this gene
AKO66_01345
AKO66_01350
248. : CP012329 Bacillus cellulasensis strain NJ-M2 Total score: 1.0 Cumulative Blast bit score: 261
AKO65_05005
AKO65_05010
AKO65_05015
AKO65_05020
transcription antiterminator LicT
Accession: ALM27397
Location: 1024964-1025797
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ALM27397
Location: 1024964-1025797
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
AKO65_05025
249. : CP011109 Bacillus pumilus strain C4 genome. Total score: 1.0 Cumulative Blast bit score: 261
transcription antiterminator LicT
Accession: ANT58472
Location: 3395899-3396732
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: ANT58472
Location: 3395899-3396732
BlastP hit with bglK
Percentage identity: 43 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
VP59_17195
PTS beta-glucoside transporter subunit IIA
Accession: ANT58471
Location: 3393666-3395573
NCBI BlastP on this gene
Accession: ANT58471
Location: 3393666-3395573
NCBI BlastP on this gene
VP59_17190
VP59_17185
250. : CP009108 Bacillus altitudinis strain GR-8 Total score: 1.0 Cumulative Blast bit score: 261
ID12_04170
ID12_04165
ID12_04160
ID12_04155
transcription antiterminator LicT
Accession: AKU30682
Location: 793084-793917
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
Accession: AKU30682
Location: 793084-793917
BlastP hit with bglK
Percentage identity: 44 %
BlastP bit score: 261
Sequence coverage: 98 %
E-value: 2e-82
NCBI BlastP on this gene
ID12_04150
PTS beta-glucoside transporter subunit IIA
Accession: AKU30681
Location: 790851-792758
NCBI BlastP on this gene
Accession: AKU30681
Location: 790851-792758
NCBI BlastP on this gene
ID12_04145
ID12_04140

Detecting sequence homology at the gene cluster level with MultiGeneBlast.
Marnix H. Medema, Rainer Breitling & Eriko Takano (2013)
Molecular Biology and Evolution , 30: 1218-1223.
Marnix H. Medema, Rainer Breitling & Eriko Takano (2013)
Molecular Biology and Evolution , 30: 1218-1223.