You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000724_00804
You are here: Home > Sequence: MGYG000000724_00804
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Gemmiger; | |||||||||||
| CAZyme ID | MGYG000000724_00804 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | Exo-poly-alpha-D-galacturonosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1606; End: 3096 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 111 | 485 | 2.6e-63 | 0.9507692307692308 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.10e-55 | 15 | 403 | 13 | 398 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 2.45e-18 | 235 | 400 | 76 | 236 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN03010 | PLN03010 | 1.02e-14 | 247 | 400 | 160 | 307 | polygalacturonase |
| PLN03003 | PLN03003 | 2.40e-13 | 243 | 403 | 137 | 291 | Probable polygalacturonase At3g15720 |
| PLN02155 | PLN02155 | 3.86e-06 | 245 | 403 | 146 | 299 | polygalacturonase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADL35584.1 | 1.50e-170 | 6 | 492 | 8 | 510 |
| AOZ97711.1 | 4.28e-170 | 6 | 492 | 9 | 513 |
| QOL34313.1 | 1.24e-141 | 17 | 472 | 19 | 481 |
| QOL33043.1 | 6.41e-139 | 17 | 472 | 19 | 481 |
| BAV04097.1 | 1.30e-108 | 11 | 492 | 36 | 488 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OLP_A | 2.21e-37 | 97 | 404 | 56 | 364 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 3JUR_A | 1.63e-34 | 85 | 482 | 27 | 420 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 2UVE_A | 2.31e-30 | 47 | 439 | 114 | 530 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 7E56_A | 3.40e-10 | 240 | 420 | 96 | 274 | ChainA, Endo-polygalacturonase [Evansstolkia leycettana] |
| 6KVE_A | 3.56e-10 | 240 | 420 | 104 | 282 | ChainA, Endo-polygalacturonase [Evansstolkia leycettana] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P15922 | 1.20e-34 | 12 | 433 | 44 | 522 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
| A7PZL3 | 4.59e-34 | 95 | 400 | 72 | 358 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| P27644 | 6.65e-21 | 239 | 408 | 16 | 186 | Polygalacturonase OS=Rhizobium radiobacter OX=358 GN=pgl PE=2 SV=1 |
| P20041 | 3.42e-16 | 101 | 442 | 72 | 429 | Polygalacturonase OS=Ralstonia solanacearum OX=305 GN=pglA PE=1 SV=1 |
| P58598 | 1.39e-14 | 101 | 393 | 74 | 381 | Polygalacturonase OS=Ralstonia solanacearum (strain GMI1000) OX=267608 GN=pglA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
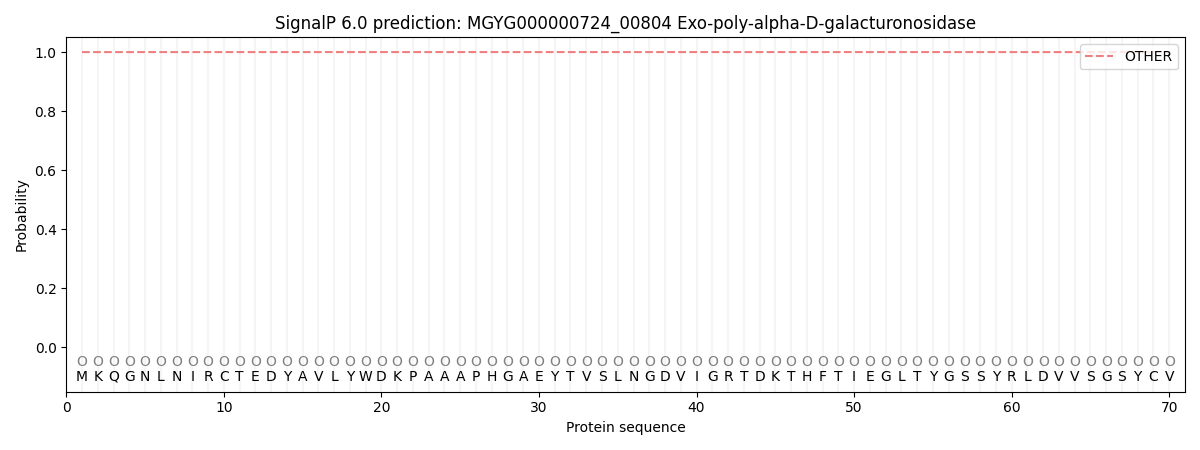
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000068 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
