You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002341_01037
You are here: Home > Sequence: MGYG000002341_01037
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Acinetobacter ursingii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Pseudomonadales; Moraxellaceae; Acinetobacter; Acinetobacter ursingii | |||||||||||
| CAZyme ID | MGYG000002341_01037 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 65450; End: 67276 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE4 | 93 | 257 | 1.1e-24 | 0.9230769230769231 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR03938 | deacetyl_PgaB | 3.17e-121 | 37 | 396 | 3 | 360 | poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB. Two well-characterized systems produce polysaccharide based on N-acetyl-D-glucosamine in straight chains with beta-1,6 linkages. These are encoded by the icaADBC operon in Staphylococcus species, where the system is designated polysaccharide intercellular adhesin (PIA), and the pgaABCD operon in Gram-negative bacteria such as E. coli. Both systems include a putative polysaccharide deacetylase. The PgaB protein, described here, contains an additional domain lacking from its Gram-positive counterpart IcaB (TIGR03933). Deacetylation by this protein appears necessary to allow export through the porin PgaA [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
| cd10964 | CE4_PgaB_5s | 1.91e-83 | 95 | 263 | 1 | 173 | N-terminal putative catalytic polysaccharide deacetylase domain of bacterial poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB, and similar proteins. This family is represented by an outer membrane lipoprotein, poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase (PgaB, EC 3.5.1.-), encoded by Escherichia coli pgaB gene from the pgaABCD (formerly ycdSRQP) operon, which affects biofilm development by promoting abiotic surface binding and intercellular adhesion. PgaB catalyzes the N-deacetylation of poly-beta-1,6-N-acetyl-D-glucosamine (PGA), a biofilm adhesin polysaccharide that stabilizes biofilms of E. coli and other bacteria. PgaB contains an N-terminal NodB homology domain with a 5-stranded beta/alpha barrel, and a C-terminal carbohydrate binding domain required for PGA N-deacetylation, which may be involved in binding to unmodified poly-beta-1,6-GlcNAc and assisting catalysis by the deacetylase domain. This family also includes several orthologs of PgaB, such as the hemin storage system HmsF protein, encoded by Yersinia pestis hmsF gene from the hmsHFRS operon, which is essential for Y. pestis biofilm formation. Like PgaB, HmsF is an outer membrane protein with an N-terminal NodB homology domain, which is likely involved in the modification of the exopolysaccharide (EPS) component of the biofilm. HmsF also has a conserved but uncharacterized C-terminal domain that is present in other HmsF-like proteins in Gram-negative bacteria. This alignment model corresponds to the N-terminal NodB homology domain. |
| PRK14582 | pgaB | 1.84e-71 | 36 | 395 | 48 | 402 | poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB. |
| PRK14581 | hmsF | 7.06e-64 | 34 | 405 | 46 | 413 | outer membrane N-deacetylase; Provisional |
| cd10918 | CE4_NodB_like_5s_6s | 9.40e-33 | 99 | 260 | 1 | 134 | Putative catalytic NodB homology domain of PgaB, IcaB, and similar proteins which consist of a deformed (beta/alpha)8 barrel fold with 5- or 6-strands. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes bacterial poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB, hemin storage system HmsF protein in gram-negative species, intercellular adhesion proteins IcaB, and many uncharacterized prokaryotic polysaccharide deacetylases. It also includes a putative polysaccharide deacetylase YxkH encoded by the Bacillus subtilis yxkH gene, which is one of six polysaccharide deacetylase gene homologs present in the Bacillus subtilis genome. Sequence comparison shows all family members contain a conserved domain similar to the catalytic NodB homology domain of rhizobial NodB-like proteins, which consists of a deformed (beta/alpha)8 barrel fold with 6 or 7 strands. However, in this family, most proteins have 5 strands and some have 6 strands. Moreover, long insertions are found in many family members, whose function remains unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQT86461.1 | 0.0 | 1 | 608 | 1 | 608 |
| QQT66978.1 | 0.0 | 1 | 608 | 1 | 608 |
| BBF77886.1 | 0.0 | 1 | 608 | 1 | 608 |
| QPF34271.1 | 2.69e-228 | 24 | 599 | 26 | 603 |
| AUC05455.1 | 2.69e-228 | 24 | 599 | 26 | 603 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4F9D_A | 1.98e-50 | 37 | 396 | 12 | 366 | Structureof Escherichia coli PgaB 42-655 in complex with nickel [Escherichia coli K-12],4F9D_B Structure of Escherichia coli PgaB 42-655 in complex with nickel [Escherichia coli K-12] |
| 5BU6_A | 4.51e-50 | 37 | 261 | 13 | 244 | Structureof BpsB deaceylase domain from Bordetella bronchiseptica [Bordetella bronchiseptica RB50],5BU6_B Structure of BpsB deaceylase domain from Bordetella bronchiseptica [Bordetella bronchiseptica RB50] |
| 4F9J_A | 4.72e-49 | 37 | 396 | 12 | 366 | Structureof Escherichia coli PgaB 42-655 in complex with iron [Escherichia coli K-12],4F9J_B Structure of Escherichia coli PgaB 42-655 in complex with iron [Escherichia coli K-12] |
| 4U10_A | 3.57e-38 | 36 | 262 | 3 | 233 | Probingthe structure and mechanism of de-N-acetylase from aggregatibacter actinomycetemcomitans [Aggregatibacter actinomycetemcomitans],4U10_B Probing the structure and mechanism of de-N-acetylase from aggregatibacter actinomycetemcomitans [Aggregatibacter actinomycetemcomitans] |
| 3VUS_A | 9.48e-38 | 37 | 293 | 8 | 265 | Escherichiacoli PgaB N-terminal domain [Escherichia coli K-12],3VUS_B Escherichia coli PgaB N-terminal domain [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8XAR3 | 1.63e-49 | 37 | 396 | 49 | 403 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Escherichia coli O157:H7 OX=83334 GN=pgaB PE=3 SV=1 |
| P75906 | 2.24e-49 | 37 | 396 | 49 | 403 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Escherichia coli (strain K12) OX=83333 GN=pgaB PE=1 SV=1 |
| Q8NUI6 | 1.51e-16 | 57 | 288 | 74 | 279 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain MW2) OX=196620 GN=icaB PE=3 SV=1 |
| Q7A349 | 1.51e-16 | 57 | 288 | 74 | 279 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain N315) OX=158879 GN=icaB PE=3 SV=1 |
| Q99QX2 | 1.51e-16 | 57 | 288 | 74 | 279 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=icaB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
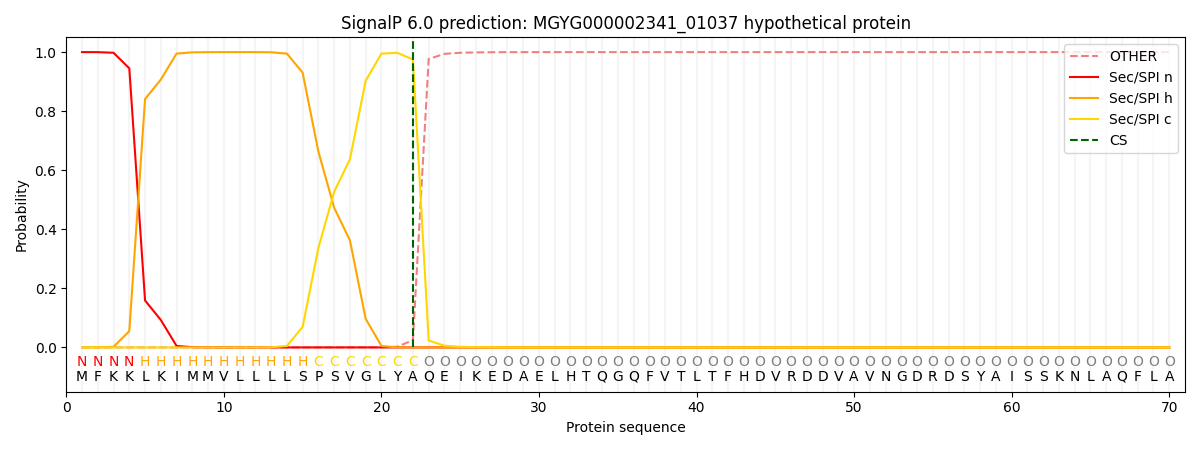
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000320 | 0.998887 | 0.000221 | 0.000197 | 0.000183 | 0.000163 |
