You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003312_03583
You are here: Home > Sequence: MGYG000003312_03583
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp900547205 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp900547205 | |||||||||||
| CAZyme ID | MGYG000003312_03583 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10379; End: 11920 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH30 | 63 | 508 | 7e-121 | 0.9822222222222222 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5520 | XynC | 1.40e-28 | 56 | 511 | 35 | 429 | O-Glycosyl hydrolase [Cell wall/membrane/envelope biogenesis]. |
| pfam02057 | Glyco_hydro_59 | 4.38e-07 | 87 | 369 | 24 | 251 | Glycosyl hydrolase family 59. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIU93337.1 | 0.0 | 1 | 513 | 13 | 525 |
| QUT40981.1 | 0.0 | 1 | 513 | 1 | 513 |
| BCA52347.1 | 2.13e-154 | 1 | 513 | 1 | 515 |
| QQA08798.1 | 8.58e-154 | 1 | 513 | 1 | 515 |
| ALJ39695.1 | 3.45e-153 | 1 | 513 | 1 | 515 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4FMV_A | 2.16e-21 | 66 | 510 | 14 | 386 | CrystalStructure Analysis of a GH30 Endoxylanase from Clostridium papyrosolvens C71 [Ruminiclostridium papyrosolvens DSM 2782] |
| 6KRL_A | 1.07e-12 | 25 | 418 | 57 | 455 | Crystalstructure of GH30 xylanase B from Talaromyces cellulolyticus expressed by Pichia pastoris [Saccharomyces uvarum],6KRN_A Crystal structure of GH30 xylanase B from Talaromyces cellulolyticus expressed by Pichia pastoris in complex with aldotriuronic acid [Saccharomyces uvarum] |
| 6IUJ_A | 3.74e-12 | 54 | 418 | 21 | 384 | ChainA, GH30 Xylanase B [Talaromyces cellulolyticus CF-2612],6IUJ_B Chain B, GH30 Xylanase B [Talaromyces cellulolyticus CF-2612] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| G5ECR8 | 1.37e-12 | 57 | 473 | 93 | 517 | Putative glucosylceramidase 3 OS=Caenorhabditis elegans OX=6239 GN=gba-3 PE=3 SV=1 |
| P17439 | 1.25e-06 | 53 | 450 | 82 | 478 | Lysosomal acid glucosylceramidase OS=Mus musculus OX=10090 GN=Gba PE=1 SV=1 |
| Q2KHZ8 | 6.81e-06 | 53 | 458 | 102 | 518 | Lysosomal acid glucosylceramidase OS=Bos taurus OX=9913 GN=GBA PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
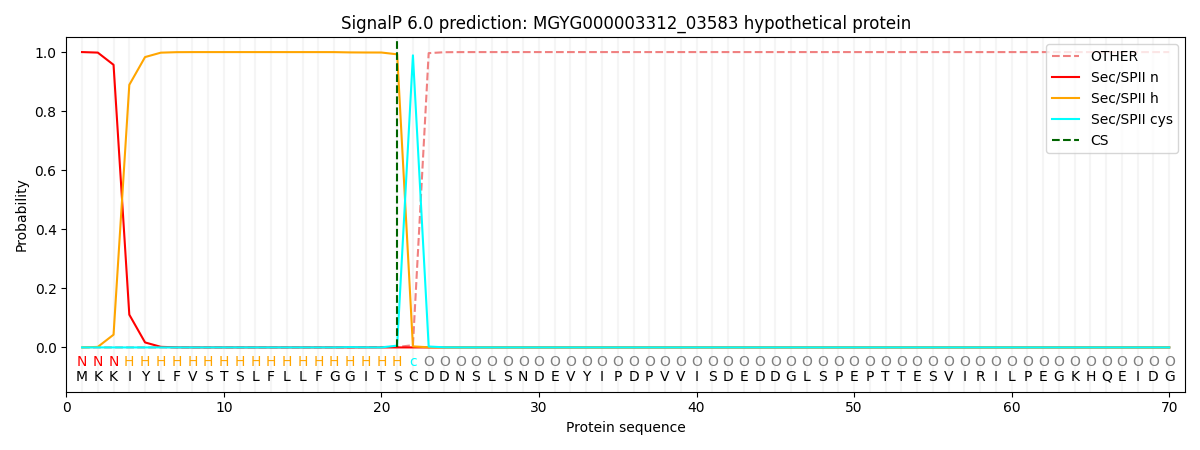
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000005 | 1.000063 | 0.000000 | 0.000000 | 0.000000 |
