You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003322_01284
You are here: Home > Sequence: MGYG000003322_01284
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Coriobacteriaceae; Collinsella; | |||||||||||
| CAZyme ID | MGYG000003322_01284 | |||||||||||
| CAZy Family | CE19 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 20; End: 985 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE19 | 63 | 302 | 1.3e-26 | 0.5933734939759037 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam12715 | Abhydrolase_7 | 1.17e-09 | 81 | 287 | 128 | 332 | Abhydrolase family. This is a family of probable bacterial abhydrolases. |
| COG0412 | DLH | 2.75e-07 | 72 | 219 | 23 | 153 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
| pfam01738 | DLH | 4.15e-07 | 92 | 209 | 24 | 127 | Dienelactone hydrolase family. |
| COG1506 | DAP2 | 2.82e-06 | 103 | 237 | 419 | 525 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| pfam00326 | Peptidase_S9 | 1.03e-04 | 103 | 209 | 10 | 96 | Prolyl oligopeptidase family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ATP54718.1 | 3.51e-219 | 1 | 317 | 400 | 716 |
| QIA34433.1 | 1.06e-217 | 1 | 317 | 400 | 716 |
| QUC03567.1 | 1.11e-159 | 3 | 316 | 396 | 709 |
| QNN25394.1 | 8.60e-20 | 1 | 318 | 58 | 358 |
| VIP10631.1 | 6.93e-19 | 3 | 298 | 77 | 344 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34973 | 4.78e-16 | 1 | 288 | 9 | 280 | Putative hydrolase YtaP OS=Bacillus subtilis (strain 168) OX=224308 GN=ytaP PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
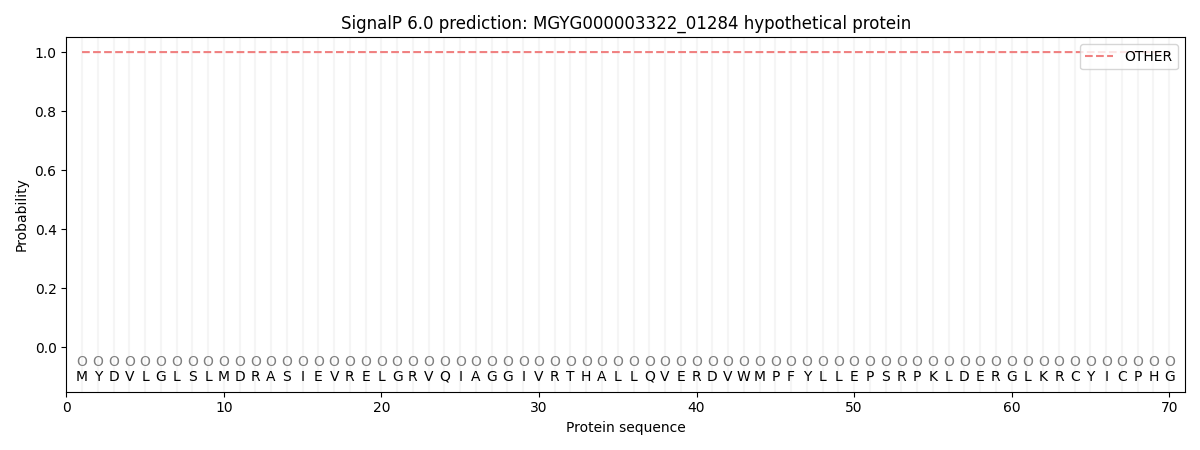
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000066 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
