

| Basic Information | |
|---|---|
| Species | Selaginella moellendorffii |
| Cazyme ID | 410831 |
| Family | GH103 |
| Protein Properties | Length: 1367 Molecular Weight: 149818 Isoelectric Point: 9.2391 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH103 | 1057 | 1353 | 0 |
| AWLEDLKEEAIQRGVGKDTVLEAFDGVKISPKIMEKDQTQPEKTLTAEDYIKTMVSQSRVSRGVQAFVDNKALLEEISAEYGVPASLLVSIWGIESNYGS YTGSWNVIQALVTLAFDSPEARRRAFFRDEVFHALFILDKAEGPSNAAGLKGSWAGAMGQCQFMPSSFRNYAVDFDRDGRKDIWSSKADIFASIANYLAE HGWEIGGKYAQKVKIPSDMDKSVYGLDIKKNVNRWLEENTITAADGSQLSGDEPVSLVAPDGPSGVAYLVWENFRVVMRYNISILYSLAVYELAEEI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1367 Download |
| MTLLNAYRPE WSPAAINSAI VTTGFVTVES RSKTQHCSPP PLSTSVEATS IQTRLLTRAS 60 STIGARLHGL SVWIEAGKTK RPLSRTCSIR RRMASRNLNY PSIAISSLKG KRTIHRRVTN 120 VDDDATTYRA SVESPDGVLG LSKAIYVFSK SDDVNAHSSR KKSYQSALVI PGSTEVAKHG 180 KTESTQSSEF CSLPQRFLVM VLYRNSKILD PVSRRDADLI KSRRWMTALN FSIFLLDNMI 240 FEQKAIHIVY LGDLDATLHS DPDAVSDSHY QLLENILGSY RHGFNGFAAS LTSEEAAHIS 300 RIPNVISVFP TQNLKLHTTN SWSFLGFDED GGKNVQSLFG DSESKSLRRK ANYGEDIIVG 360 VIDTGTNIAR VQELCGRFNG SNIAEMEGHL RARRELHFLA LQQEAHRCKV FRAGRKNQHG 420 DIFSRDANGH GTHAASIAGG RFVENVSWSG YASSTLSGGA SNSRIAAYKV CWARQQAFDC 480 TSESVLAAFE AGIHDRVGAF SVSLGTNSSD YFSDPIAIGC IVVVAAAGNN YSFGTVANAA 540 PWIITVAAST ISRSFTSRGS LGNNQSFAGL SLTRHTLGRR WYPLVLSLDA GIPNSSFTTS 600 ELERCVSGSL DPKKVRGKIV TCLDTLPLGG GARQAGIIFC NRGPQSARPF EHALPASHVN 660 RAEANAIQSY INSTSPLLNT KPAPMMGVFS SQGPNPVTPD IFKPDITTPG VDILAAYSPF 720 ANNRGISYCV WNFHGLRKSD FFSRLQKVFS LITLESRSKT RHCFPPPLST SVEATSIQAR 780 LLIRASSTMP RRPTSRTFGL RQERRYISTF ANCATRRRMA SSDLNYPSIA TSSLRGKRIV 840 HRRVTNVDDD ATTYRSSVEA PDGVLVSVVP PVLEFERRGE TIAFKVRFQA VKASNQSYAF 900 GTLEQWKGCL IPAMSAMAFA SPGLRFHSRD GMFAKLDPSC RILPPFSLVS SRCRAFPRKL 960 ALAMCSSQQQ SRVSGLWNLG QIERCLEAIH CALDPKLAAV LVVVACSSFR REGHSYTSQA 1020 PVFCSIASEQ PAIRAQIHVK INQIPVEMST PESDISAWLE DLKEEAIQRG VGKDTVLEAF 1080 DGVKISPKIM EKDQTQPEKT LTAEDYIKTM VSQSRVSRGV QAFVDNKALL EEISAEYGVP 1140 ASLLVSIWGI ESNYGSYTGS WNVIQALVTL AFDSPEARRR AFFRDEVFHA LFILDKAEGP 1200 SNAAGLKGSW AGAMGQCQFM PSSFRNYAVD FDRDGRKDIW SSKADIFASI ANYLAEHGWE 1260 IGGKYAQKVK IPSDMDKSVY GLDIKKNVNR WLEENTITAA DGSQLSGDEP VSLVAPDGPS 1320 GVAYLVWENF RVVMRYNISI LYSLAVYELA EEITAGMTAL ESKANE* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK10760 | PRK10760 | 2.0e-40 | 1106 | 1353 | 255 | + murein hydrolase B; Provisional | ||
| TIGR02282 | MltB | 2.0e-63 | 1062 | 1353 | 294 | + lytic murein transglycosylase B. This family consists of lytic murein transglycosylases (murein hydrolases) in the family of MltB, which is a membrane-bound lipoprotein in Escherichia coli. The N-terminal lipoprotein modification motif is conserved in about half the members of this family. The term Slt35 describes a naturally occurring soluble fragment of MltB. Members of this family never contain the putative peptidoglycan binding domain described by pfam01471, which is associated with several classes of bacterial cell wall lytic enzymes [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| COG2951 | MltB | 8.0e-94 | 1052 | 1353 | 305 | + Membrane-bound lytic murein transglycosylase B [Cell envelope biogenesis, outer membrane] | ||
| TIGR02283 | MltB_2 | 6.0e-101 | 1054 | 1354 | 305 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| pfam13406 | SLT_2 | 1.0e-123 | 1056 | 1351 | 296 | + Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002266135.1 | 0.007 | 3 | 138 | 592 | 733 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002266135.1 | 0 | 247 | 903 | 58 | 773 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002299062.1 | 0.002 | 3 | 150 | 574 | 727 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002299062.1 | 0 | 239 | 908 | 24 | 759 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002306266.1 | 0 | 247 | 903 | 11 | 726 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3vta_B | 0 | 318 | 886 | 1 | 587 | A Chain A, Crystal Structure Of Cucumisin, A Subtilisin-Like Endoprotease From Cucumis Melo L |
| PDB | 3vta_A | 0 | 318 | 886 | 1 | 587 | A Chain A, Crystal Structure Of Cucumisin, A Subtilisin-Like Endoprotease From Cucumis Melo L |
| PDB | 4anr_A | 3.00018e-42 | 1070 | 1353 | 37 | 316 | A Chain A, Crystal Structure Of Soluble Lytic Transglycosylase Sltb1 From Pseudomonas Aeruginosa |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EX384096 | 288 | 1057 | 1343 | 0 |
| EX382166 | 268 | 1036 | 1302 | 0 |
| GO366029 | 265 | 1036 | 1299 | 0 |
| EX384473 | 239 | 1120 | 1357 | 0 |
| HO416716 | 315 | 1044 | 1356 | 0 |
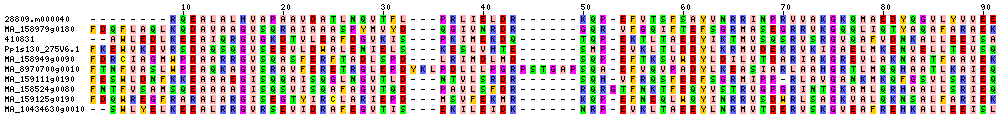
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |