

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s130_275V6.1 |
| Family | GH103 |
| Protein Properties | Length: 531 Molecular Weight: 58704 Isoelectric Point: 6.5238 |
| Chromosome | Chromosome/Scaffold: 130 Start: 1041771 End: 1044543 |
| Description | |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH103 | 212 | 517 | 0 |
| FKEWVKDVRSDAQSQGVSEEVLDWALENIELSKESLVHTESMPEVKLTLDDYLKRMVDEKRVKIGAELMKENVELLTEVSQKFGVPTQFLVAIWGIETNY GSYMGNWDIIQSLATMSFTEKPASRSNYFRGELIEALLMLQKGTADRGRGPEGKTRLRGSWAGAMGQCQFMPTSFRDYALDYDGDGHKDIWESKADVFAS IANYFKEHGWKEGGPIFQKVQVPRKLDESLVGLGVVKSVLDWEHKHGVHLNPGKAPPLPPDEMASLLMPNGPKGNAYLVCSNFRVILRYNSSSLYGFSVC ELARLI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 531 Download |
| MALFSLSSAL LSPYHGSVAE LKEPSALKIL YPYQFPSLGR FRRGGIALPA FGGLGWRSVA 60 EQTIESVHLH SHRCFSFPEK DSNSTTELQG PEFALEDECT DGAMSSGRKM IMLFLSKSVI 120 IGVLAFTMAA VAAASRNRRT GLVAEPADSG QVYVLNGKPV VSISNKFNEA EPRISSREKL 180 SRSLCFASTA LTANDGLTKL QEKAIDEAMQ KFKEWVKDVR SDAQSQGVSE EVLDWALENI 240 ELSKESLVHT ESMPEVKLTL DDYLKRMVDE KRVKIGAELM KENVELLTEV SQKFGVPTQF 300 LVAIWGIETN YGSYMGNWDI IQSLATMSFT EKPASRSNYF RGELIEALLM LQKGTADRGR 360 GPEGKTRLRG SWAGAMGQCQ FMPTSFRDYA LDYDGDGHKD IWESKADVFA SIANYFKEHG 420 WKEGGPIFQK VQVPRKLDES LVGLGVVKSV LDWEHKHGVH LNPGKAPPLP PDEMASLLMP 480 NGPKGNAYLV CSNFRVILRY NSSSLYGFSV CELARLIEER SSNPYPTPAL * 540 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK10760 | PRK10760 | 6.0e-41 | 263 | 514 | 277 | + murein hydrolase B; Provisional | ||
| TIGR02282 | MltB | 4.0e-63 | 223 | 514 | 298 | + lytic murein transglycosylase B. This family consists of lytic murein transglycosylases (murein hydrolases) in the family of MltB, which is a membrane-bound lipoprotein in Escherichia coli. The N-terminal lipoprotein modification motif is conserved in about half the members of this family. The term Slt35 describes a naturally occurring soluble fragment of MltB. Members of this family never contain the putative peptidoglycan binding domain described by pfam01471, which is associated with several classes of bacterial cell wall lytic enzymes [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| COG2951 | MltB | 1.0e-90 | 176 | 517 | 343 | + Membrane-bound lytic murein transglycosylase B [Cell envelope biogenesis, outer membrane] | ||
| TIGR02283 | MltB_2 | 4.0e-100 | 212 | 514 | 304 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| pfam13406 | SLT_2 | 2.0e-126 | 212 | 515 | 304 | + Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAM76692.1 | 0 | 201 | 517 | 27 | 327 | Membrane-bound lytic murein transglycosylase B [Magnetospirillum gryphiswaldense MSR-1] |
| RefSeq | NP_797255.1 | 0 | 212 | 517 | 24 | 321 | putative lytic murein transglycosylase [Vibrio parahaemolyticus RIMD 2210633] |
| RefSeq | XP_001770826.1 | 0 | 1 | 530 | 1 | 530 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | ZP_01043740.1 | 0 | 210 | 517 | 43 | 339 | Membrane-bound lytic murein transglycosylase [Idiomarina baltica OS145] |
| RefSeq | ZP_01312886.1 | 0 | 211 | 520 | 41 | 340 | Lytic murein transglycosylase [Desulfuromonas acetoxidans DSM 684] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 4anr_A | 4.99983e-42 | 244 | 514 | 53 | 313 | A Chain A, Crystal Structure Of Soluble Lytic Transglycosylase Sltb1 From Pseudomonas Aeruginosa |
| PDB | 1ltm_A | 5e-34 | 263 | 517 | 76 | 314 | A Chain A, Accelerated X-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using W |
| PDB | 1qut_A | 6e-34 | 263 | 517 | 78 | 316 | A Chain A, Accelerated X-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using W |
| PDB | 1qus_A | 6e-34 | 263 | 517 | 78 | 316 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdt_A | 6e-34 | 263 | 517 | 78 | 316 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| Transmembrane Domains | |||||
|---|---|---|---|---|---|
 | |||||
| Start | End | ||||
| 113 | 135 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| BY946922 | 251 | 13 | 263 | 0 |
| BJ604140 | 216 | 316 | 531 | 0 |
| BY949902 | 205 | 13 | 217 | 0 |
| BY957613 | 199 | 13 | 211 | 0 |
| BY949902 | 28 | 212 | 239 | 0.005 |
| Orthologous Group | |||||
|---|---|---|---|---|---|
| Species | ID | ||||
| Picea abies | MA_158979g0180 | MA_158949g0090 | MA_8970700g0010 | MA_159111g0190 | MA_158524g0080 |
| MA_159125g0190 | MA_10434630g0010 | ||||
| Ricinus communis | 28809.m000040 | ||||
| Selaginella moellendorffii | 410831 | ||||
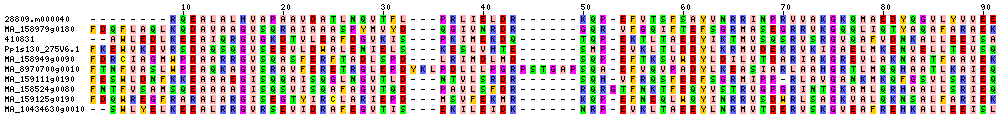
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |