

| Basic Information | |
|---|---|
| Species | Eucalyptus grandis |
| Cazyme ID | Eucgr.A02320.1 |
| Family | GT1 |
| Protein Properties | Length: 463 Molecular Weight: 49958 Isoelectric Point: 6.0171 |
| Chromosome | Chromosome/Scaffold: 1 Start: 33942425 End: 33943813 |
| Description | UDP-glycosyltransferase 73B4 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 251 | 440 | 2.3e-38 |
| CMEWLDKQAPSSVIYVSFGTTTVLCDEQIREIAMGLERSGQKFIWVLREADKADIFGGGGEARKIELPQGFEEMAAARDLGVVVRDWAPQLEILGHPATG GFLSHCGWNSCMESISMGVPILAWPMHSDQPLNAVLITQVLKIGLILKDWMSPDDVVKSSAIEGAVKTLMASEEGEEARKRAAELGAAVR | |||
| Full Sequence |
|---|
| Protein Sequence Length: 463 Download |
| MVPLPAQGHL NQLLQLAHLV ASCGVPVHYV GSSSHNRQAK LRSHVQPSSV PSDICFHDFP 60 TPPIPSPAPS PNSPYKFPSH LQPAFDATSS LGGPVASLVR SLSSVSRRVV VVYDSLMASV 120 VQDAASVPNA EIYVFQPPSA FTLCWFTWES IGMAPPPEVE VDLAEFISSK GFPSLLDSYP 180 NEFLKFMESQ KECQKSNSGC IHNTCRVVEG RFVDLFASAT ANGTAIKHWA IGPLNPVKVP 240 EKSSRRSAHG CMEWLDKQAP SSVIYVSFGT TTVLCDEQIR EIAMGLERSG QKFIWVLREA 300 DKADIFGGGG EARKIELPQG FEEMAAARDL GVVVRDWAPQ LEILGHPATG GFLSHCGWNS 360 CMESISMGVP ILAWPMHSDQ PLNAVLITQV LKIGLILKDW MSPDDVVKSS AIEGAVKTLM 420 ASEEGEEARK RAAELGAAVR GSMDEGGVSR AELDSFIAHV SR* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02554 | PLN02554 | 5.0e-43 | 158 | 460 | 330 | + UDP-glycosyltransferase family protein | ||
| PLN02448 | PLN02448 | 8.0e-46 | 229 | 461 | 240 | + UDP-glycosyltransferase family protein | ||
| PLN00164 | PLN00164 | 6.0e-46 | 185 | 453 | 284 | + glucosyltransferase; Provisional | ||
| PLN02863 | PLN02863 | 4.0e-47 | 3 | 462 | 499 | + UDP-glucoronosyl/UDP-glucosyl transferase family protein | ||
| PLN03007 | PLN03007 | 3.0e-56 | 182 | 461 | 291 | + UDP-glucosyltransferase family protein | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002284381.1 | 0 | 1 | 462 | 21 | 473 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002308926.1 | 0 | 1 | 461 | 25 | 472 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002308931.1 | 0 | 1 | 461 | 25 | 472 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002322723.1 | 0 | 1 | 462 | 1 | 452 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002532137.1 | 0 | 1 | 461 | 27 | 455 | UDP-glucosyltransferase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2vg8_A | 0 | 1 | 451 | 11 | 459 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2vch_A | 0 | 1 | 451 | 11 | 459 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2vce_A | 0 | 1 | 451 | 11 | 459 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 2pq6_A | 4.00001e-41 | 1 | 460 | 13 | 477 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2acv_B | 2e-37 | 195 | 461 | 210 | 462 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EG631252 | 467 | 1 | 463 | 0 |
| ES588947 | 212 | 75 | 286 | 0 |
| FY800511 | 166 | 191 | 356 | 0 |
| GR998914 | 285 | 179 | 463 | 0 |
| FY800511 | 35 | 151 | 184 | 2.1 |
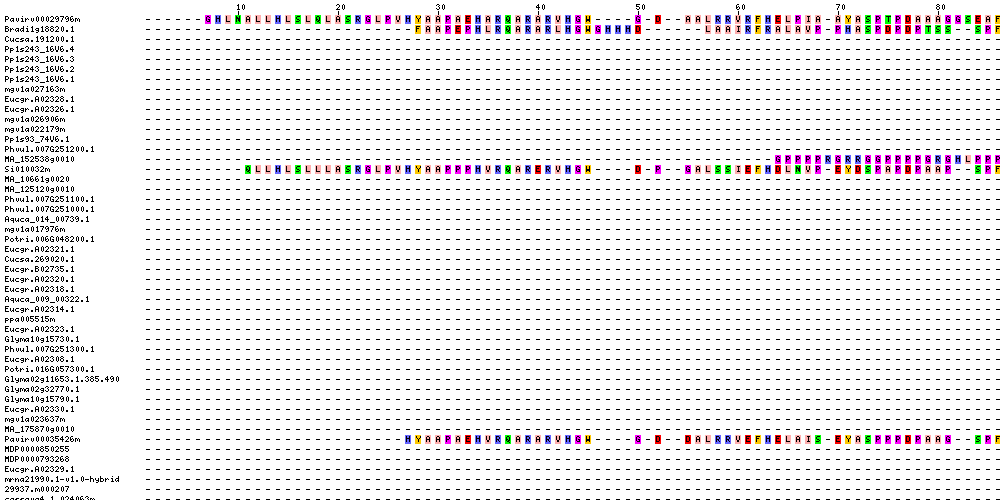
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |