

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s64_78V6.1 |
| Family | GH13 |
| Protein Properties | Length: 734 Molecular Weight: 84470.4 Isoelectric Point: 6.5658 |
| Chromosome | Chromosome/Scaffold: 64 Start: 819453 End: 825817 |
| Description | starch branching enzyme 2.1 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH13 | 246 | 570 | 1.9e-26 |
| LPRIKANNYNTIQLMAIMEHAYYGCFGYHVTNFFAVSSRSGTPEDLKYLIDQAHSMGLRVLMDVVHSHASTNAVDGLAGYDLGQPAQESYFHTGARGYHK LWDSRLFNYGSWEVQRFLLSNLRWWMDEYMFDGFRFDGITSMLYHHHGLNMRFSGNYYEYFSEATDVEAVMYLMLANDLVHKMYPDATVIAEDVSGFPTL CRPVSEGGVGFDYRLAMGIPDKWMEYLKVKEEEDWSMGDIVHTLTNRRYKEPCVAYSESHDQSMVGDKSYAFLLMDKEMYFSMLATQPSNPIVDRGIALH KMIHFITMALGGEGYLNFMGNEFGH | |||
| Full Sequence |
|---|
| Protein Sequence Length: 734 Download |
| MWGVGGLTRR FSFKASCPTV KASDLEADKP DNSADKPQPW KDMNGMQNLG VLELDPYLAP 60 HRIHLRSRVR EFMKRKMEIE KHEGSLEDFA EGYKKFGFTR EGNCIVYREW APAAAAAQLI 120 GDFNNWDGRK HNMERDKFGV WSIRLPDVRG VSAIPHGSKV RIRIKKGDDT WVDRIPAWIK 180 YAVVDPTVFA ADYDGVYWDP PLRERYHFKH ARPQKPSAPL IYEAHVGMSS KEPRVSSYRE 240 FADEVLPRIK ANNYNTIQLM AIMEHAYYGC FGYHVTNFFA VSSRSGTPED LKYLIDQAHS 300 MGLRVLMDVV HSHASTNAVD GLAGYDLGQP AQESYFHTGA RGYHKLWDSR LFNYGSWEVQ 360 RFLLSNLRWW MDEYMFDGFR FDGITSMLYH HHGLNMRFSG NYYEYFSEAT DVEAVMYLML 420 ANDLVHKMYP DATVIAEDVS GFPTLCRPVS EGGVGFDYRL AMGIPDKWME YLKVKEEEDW 480 SMGDIVHTLT NRRYKEPCVA YSESHDQSMV GDKSYAFLLM DKEMYFSMLA TQPSNPIVDR 540 GIALHKMIHF ITMALGGEGY LNFMGNEFGH PEWIDFPRQG NNWSFDKCRR LWDLADRDDL 600 RYKFMNNFNR AMIGLGESFQ FVGSSKQYIS NKSETERVIA FERGDLVFVF NFHSTNTYPE 660 LKVGCEIPGN YRICLDSDAA EFGGHSRVDH EVNHCTNPEG EPGKPETGYN NRPHSFMVMS 720 PSRSCQVYHK VSE* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02960 | PLN02960 | 2.0e-12 | 6 | 148 | 161 | + alpha-amylase | ||
| PLN03244 | PLN03244 | 6.0e-134 | 151 | 728 | 583 | + alpha-amylase; Provisional | ||
| PLN02960 | PLN02960 | 3.0e-176 | 151 | 728 | 581 | + alpha-amylase | ||
| PLN02447 | PLN02447 | 0 | 18 | 733 | 716 | + 1,4-alpha-glucan-branching enzyme | ||
| cd11321 | AmyAc_bac_euk_BE | 0 | 204 | 610 | 407 | + Alpha amylase catalytic domain found in bacterial and eukaryotic branching enzymes. Branching enzymes (BEs) catalyze the formation of alpha-1,6 branch points in either glycogen or starch by cleavage of the alpha-1,4 glucosidic linkage yielding a non-reducing end oligosaccharide chain, and subsequent attachment to the alpha-1,6 position. By increasing the number of non-reducing ends, glycogen is more reactive to synthesis and digestion as well as being more soluble. This group includes bacterial and eukaryotic proteins. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| GO:0043169 | cation binding |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAA70038.1 | 0 | 39 | 733 | 11 | 705 | 1,4-alpha-glucan branching enzyme [Solanum tuberosum] |
| EMBL | CBI18866.1 | 0 | 47 | 733 | 94 | 780 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_001754772.1 | 0 | 46 | 733 | 1 | 688 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001762321.1 | 0 | 46 | 727 | 1 | 682 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001763855.1 | 0 | 46 | 733 | 1 | 688 | predicted protein [Physcomitrella patens subsp. patens] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3aml_A | 0 | 46 | 733 | 8 | 695 | A Chain A, Structure Of The Starch Branching Enzyme I (Bei) From Oryza Sativa L |
| PDB | 3amk_A | 0 | 46 | 733 | 8 | 695 | A Chain A, Structure Of The Starch Branching Enzyme I (Bei) From Oryza Sativa L |
| PDB | 3vu2_B | 0 | 46 | 733 | 8 | 695 | A Chain A, Structure Of The Starch Branching Enzyme I (bei) Complexed With Maltopentaose From Oryza Sativa L |
| PDB | 3vu2_A | 0 | 46 | 733 | 8 | 695 | A Chain A, Structure Of The Starch Branching Enzyme I (bei) Complexed With Maltopentaose From Oryza Sativa L |
| PDB | 1m7x_D | 5.00264e-43 | 110 | 684 | 32 | 577 | A Chain A, The X-Ray Crystallographic Structure Of Branching Enzyme |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| starch biosynthesis | RXN-7710 | EC-2.4.1.18 | 1,4-α-glucan branching enzyme |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO619167 | 600 | 135 | 733 | 0 |
| HO794536 | 690 | 51 | 728 | 0 |
| HO777638 | 636 | 105 | 728 | 0 |
| HO458123 | 404 | 339 | 731 | 0 |
| HO777638 | 47 | 54 | 100 | 0.025 |
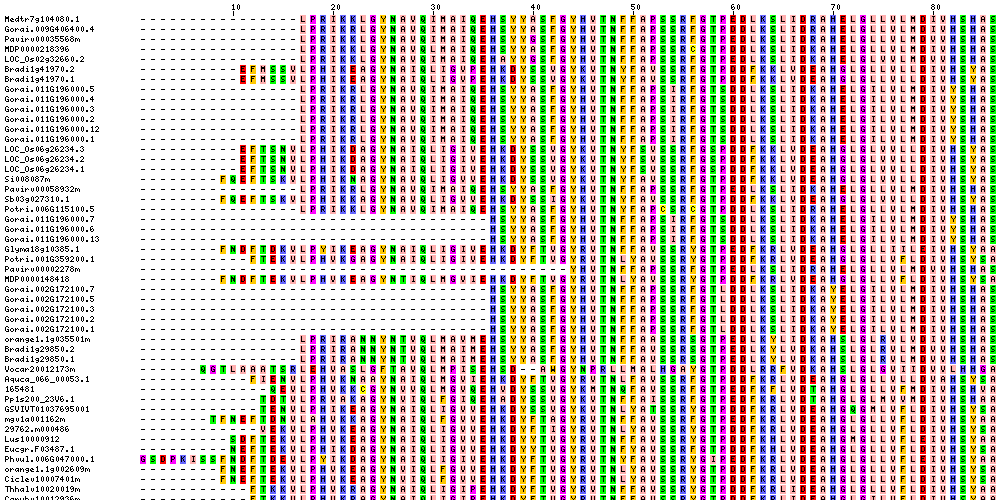
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |