

| Basic Information | |
|---|---|
| Species | Setaria italica |
| Cazyme ID | Si016917m |
| Family | GT1 |
| Protein Properties | Length: 520 Molecular Weight: 54069.4 Isoelectric Point: 5.1137 |
| Chromosome | Chromosome/Scaffold: 1 Start: 5481093 End: 5482960 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 43 | 464 | 2.2e-36 |
| AKGLLVTFSSVSSVGAKLAASAGVSSGGDGVRVGRGRVRFEFLEDGDPAGPDLDDLMRHLESAGPPAFAALLRRQASEGRPVACVVANPFMPWTIGVAAG AGIPSAVLWVQSCAVFSLYYHHVHGLVEFPPEDDAEARFSLPGLPEMTVADVPSFLLPSNPYKLLAGAIVEQFRTIGQASWVLVNSFTELESGVAAALRG VTPRPPELIPVGPLVEAVGWQDGDGDGGGDEVRGDLMKAAEECVGWLDLHPPRSVVYVSVGSVVVLSPAEVAEMAHGLASTGRPFLWVVRPDTQPHLPPG FPASVAAGRGAVVAWSPQERVLAHPSTACFLTHCGWNSTLETVAAGVPVVAFPQWGDQCTDARFLVEELGMGVRLRAGSPELQREAVREAVEDAVAGPRA EAMRASARRWSEAARRAVGPGG | |||
| Full Sequence |
|---|
| Protein Sequence Length: 520 Download |
| MGEEAAVPVA AAAASAPPHL LLICFPAQGH VNPMLRLAKR VAAKGLLVTF SSVSSVGAKL 60 AASAGVSSGG DGVRVGRGRV RFEFLEDGDP AGPDLDDLMR HLESAGPPAF AALLRRQASE 120 GRPVACVVAN PFMPWTIGVA AGAGIPSAVL WVQSCAVFSL YYHHVHGLVE FPPEDDAEAR 180 FSLPGLPEMT VADVPSFLLP SNPYKLLAGA IVEQFRTIGQ ASWVLVNSFT ELESGVAAAL 240 RGVTPRPPEL IPVGPLVEAV GWQDGDGDGG GDEVRGDLMK AAEECVGWLD LHPPRSVVYV 300 SVGSVVVLSP AEVAEMAHGL ASTGRPFLWV VRPDTQPHLP PGFPASVAAG RGAVVAWSPQ 360 ERVLAHPSTA CFLTHCGWNS TLETVAAGVP VVAFPQWGDQ CTDARFLVEE LGMGVRLRAG 420 SPELQREAVR EAVEDAVAGP RAEAMRASAR RWSEAARRAV GPGGSSDAHV QAFVDEVARR 480 AACGGAAGAA KARAQESSED ASSLVGVAER QERVEVAVP* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02152 | PLN02152 | 2.0e-49 | 19 | 477 | 476 | + indole-3-acetate beta-glucosyltransferase | ||
| PLN02173 | PLN02173 | 2.0e-55 | 19 | 419 | 403 | + UDP-glucosyl transferase family protein | ||
| PLN02210 | PLN02210 | 3.0e-63 | 120 | 477 | 363 | + UDP-glucosyl transferase | ||
| PLN02555 | PLN02555 | 3.0e-115 | 19 | 477 | 472 | + limonoid glucosyltransferase | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACD03236.1 | 0 | 1 | 504 | 1 | 492 | UDP-glycosyltransferase UGT84C2 [Avena strigosa] |
| RefSeq | NP_001147693.1 | 0 | 1 | 484 | 1 | 477 | LOC100281303 [Zea mays] |
| RefSeq | NP_001151310.1 | 0 | 1 | 481 | 1 | 478 | limonoid UDP-glucosyltransferase [Zea mays] |
| RefSeq | XP_002453431.1 | 0 | 19 | 484 | 25 | 478 | hypothetical protein SORBIDRAFT_04g005960 [Sorghum bicolor] |
| RefSeq | XP_002455167.1 | 0 | 17 | 484 | 16 | 472 | hypothetical protein SORBIDRAFT_03g005340 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2pq6_A | 0 | 14 | 480 | 5 | 480 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2c9z_A | 1.4013e-45 | 12 | 479 | 2 | 452 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2c1z_A | 1.4013e-45 | 12 | 479 | 2 | 452 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2c1x_A | 1.4013e-45 | 12 | 479 | 2 | 452 | A Chain A, Structure And Activity Of A Flavonoid 3-O Glucosyltransferase Reveals The Basis For Plant Natural Product Modification |
| PDB | 2vg8_A | 2.94273e-44 | 15 | 466 | 4 | 456 | A Chain A, Structure And Activity Of A Flavonoid 3-O Glucosyltransferase Reveals The Basis For Plant Natural Product Modification |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| cinnamate esters biosynthesis | CINNAMATE-GLUCOSYLTRANSFERASE-RXN | EC-2.4.1.177 | cinnamate β-D-glucosyltransferase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
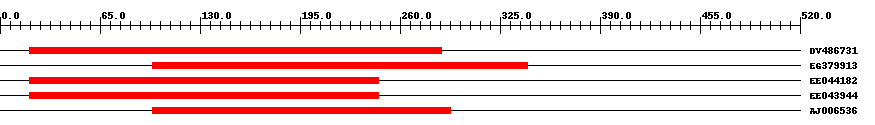 | ||||
| Hit | Length | Start | End | EValue |
| DV486731 | 269 | 19 | 287 | 0 |
| EG379913 | 245 | 99 | 343 | 0 |
| EE044182 | 228 | 19 | 246 | 0 |
| EE043944 | 228 | 19 | 246 | 0 |
| AJ006536 | 195 | 99 | 293 | 0 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |