

| Basic Information | |
|---|---|
| Species | Prunus persica |
| Cazyme ID | ppa018465m |
| Family | AA1 |
| Protein Properties | Length: 396 Molecular Weight: 43188.3 Isoelectric Point: 9.8045 |
| Chromosome | Chromosome/Scaffold: 8 Start: 11998027 End: 12000842 |
| Description | laccase 17 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA1 | 98 | 362 | 4.3e-39 |
| EANIFLALNDELFFVIAGHNLIVVEIDAVYTKSFTSQAILIVPGQTTNVLVQANQAPSRYFMAARPFMDAPVSIDNKTATRILQYKGISNTAQPVLPQLP ALINTAFALSFNAKLRSLNTAQSRPVLGISQCTTSRNGTQLTASLNNITFLMPQIGLLQAHYFYTKGVFTTDFPDCPPTPFNYTGALLTANLGTKLGTRL SKIAFNSSVELVLQDTNLLTVDGTGNFNLKKDPAKYNLVDPPERNTVGVPTGGWGAIRFRADNPG | |||
| Full Sequence |
|---|
| Protein Sequence Length: 396 Download |
| LEGDTHLVNV TNHAQYNMSI HWHGLKQYRN GRADGPAYIT QCPIKTGHSY TYNITITGQR 60 GTLWWHAHIF WLRATVYGAI VILPKQGTGF PFPQPYKEAN IFLALNDELF FVIAGHNLIV 120 VEIDAVYTKS FTSQAILIVP GQTTNVLVQA NQAPSRYFMA ARPFMDAPVS IDNKTATRIL 180 QYKGISNTAQ PVLPQLPALI NTAFALSFNA KLRSLNTAQS RPVLGISQCT TSRNGTQLTA 240 SLNNITFLMP QIGLLQAHYF YTKGVFTTDF PDCPPTPFNY TGALLTANLG TKLGTRLSKI 300 AFNSSVELVL QDTNLLTVDG TGNFNLKKDP AKYNLVDPPE RNTVGVPTGG WGAIRFRADN 360 PGKLDGVLSS FSASPSRPHA SADSPLPAQP SPQKL* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR03388 | ascorbase | 8.0e-14 | 2 | 92 | 94 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| TIGR03388 | ascorbase | 2.0e-27 | 109 | 362 | 288 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| pfam07732 | Cu-oxidase_3 | 2.0e-36 | 2 | 87 | 88 | + Multicopper oxidase. This entry contains many divergent copper oxidase-like domains that are not recognised by the pfam00394 model. | ||
| TIGR03389 | laccase | 2.0e-56 | 2 | 104 | 103 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| TIGR03389 | laccase | 6.0e-112 | 104 | 362 | 293 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005507 | copper ion binding |
| GO:0016491 | oxidoreductase activity |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI17500.1 | 0 | 2 | 362 | 47 | 500 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002266464.1 | 0 | 2 | 362 | 68 | 521 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002305436.1 | 0 | 2 | 362 | 45 | 498 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002310245.1 | 0 | 2 | 362 | 67 | 520 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002313847.1 | 0 | 2 | 362 | 66 | 519 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 4a2h_A | 0.00000000000003 | 9 | 69 | 51 | 112 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 4a2g_A | 0.00000000000003 | 9 | 69 | 51 | 112 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 4a2e_A | 0.00000000000003 | 9 | 69 | 51 | 112 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 |
| PDB | 4a2d_A | 0.00000000000003 | 9 | 69 | 51 | 112 | A Chain A, Coriolopsis Gallica Laccase T2 Copper Depleted At Ph 4.5 |
| PDB | 4a2f_A | 0.00000000000003 | 9 | 69 | 51 | 112 | A Chain A, Coriolopsis Gallica Laccase Collected At 12.65 Kev |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EX130854 | 248 | 145 | 362 | 0 |
| GO984408 | 256 | 123 | 348 | 0 |
| FC458441 | 259 | 104 | 332 | 0 |
| CF450579 | 249 | 13 | 198 | 0 |
| EY176451 | 233 | 104 | 320 | 0 |
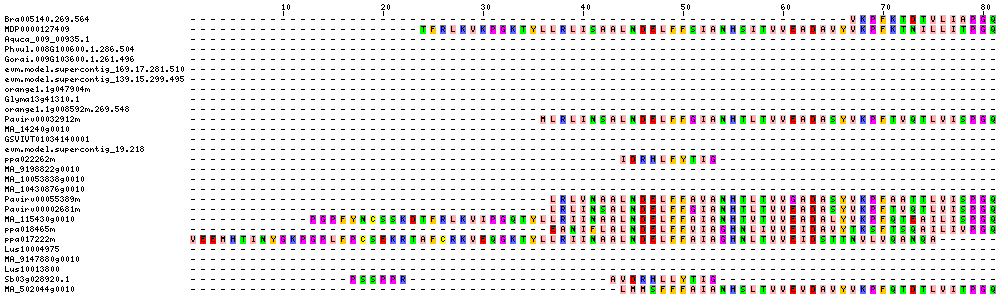
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |