

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.001G043100.2 |
| Family | GH17 |
| Protein Properties | Length: 282 Molecular Weight: 30490.8 Isoelectric Point: 7.0905 |
| Chromosome | Chromosome/Scaffold: 01 Start: 4089003 End: 4092393 |
| Description | Glycosyl hydrolase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 30 | 138 | 4e-35 |
| GINYGRIANNILSPDDVVTLLRAAKIKNVRIYDFDQSVLKAFSGTGLELVVGLPNENLRDVSANADHAMNWVKDNVLAYLPDTHIRGIAIGNEVLGSSDE FSGFLLGAI | |||
| GH17 | 137 | 240 | 1e-35 |
| AIDAAYAALEDAGYGKMEVIVTETGWASHGDENEAAATTNNARTYNYNLRKRLAKMKGTPLRPKSVLKVYVFAIFNENLKPGPTSERNFGLFKPDGSISY DIGF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 282 Download |
| MKSFGVHLFF LLFLAAIASV TVQAFTGTYG INYGRIANNI LSPDDVVTLL RAAKIKNVRI 60 YDFDQSVLKA FSGTGLELVV GLPNENLRDV SANADHAMNW VKDNVLAYLP DTHIRGIAIG 120 NEVLGSSDEF SGFLLGAIDA AYAALEDAGY GKMEVIVTET GWASHGDENE AAATTNNART 180 YNYNLRKRLA KMKGTPLRPK SVLKVYVFAI FNENLKPGPT SERNFGLFKP DGSISYDIGF 240 HGFKSSSADS SLLSLKEIRA SSWSGSYSII LTMATALLLV L* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00332 | Glyco_hydro_17 | 1.0e-15 | 153 | 240 | 88 | + Glycosyl hydrolases family 17. | ||
| pfam00332 | Glyco_hydro_17 | 8.0e-29 | 30 | 129 | 100 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABI49503.1 | 0 | 18 | 260 | 447 | 743 | Glycosyl hydrolases family 17 protein [Solanum demissum] |
| EMBL | CBI28222.1 | 0 | 6 | 281 | 11 | 339 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002282272.1 | 0 | 6 | 146 | 11 | 150 | PREDICTED: similar to beta-1,3-glucanase 1 [Vitis vinifera] |
| RefSeq | XP_002282272.1 | 0 | 129 | 281 | 240 | 391 | PREDICTED: similar to beta-1,3-glucanase 1 [Vitis vinifera] |
| RefSeq | XP_002308978.1 | 0 | 129 | 248 | 217 | 336 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1ghs_B | 9e-21 | 138 | 240 | 210 | 306 | A Chain A, The Three-Dimensional Structures Of Two Plant Beta-Glucan Endohydrolases With Distinct Substrate Specificities |
| PDB | 1ghs_B | 4e-16 | 30 | 129 | 2 | 101 | A Chain A, The Three-Dimensional Structures Of Two Plant Beta-Glucan Endohydrolases With Distinct Substrate Specificities |
| PDB | 1ghs_A | 9e-21 | 138 | 240 | 210 | 306 | A Chain A, The Three-Dimensional Structures Of Two Plant Beta-Glucan Endohydrolases With Distinct Substrate Specificities |
| PDB | 1ghs_A | 4e-16 | 30 | 129 | 2 | 101 | A Chain A, The Three-Dimensional Structures Of Two Plant Beta-Glucan Endohydrolases With Distinct Substrate Specificities |
| PDB | 2cyg_A | 1e-20 | 30 | 237 | 2 | 202 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| Transmembrane Domains | ||||
|---|---|---|---|---|
 | ||||
| Start | End | |||
| 5 | 27 | |||
| Signal Peptide | ||||
|---|---|---|---|---|
 | ||||
| Cleavage Site | ||||
| 24 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| CO118527 | 103 | 31 | 133 | 0 |
| CO120843 | 133 | 1 | 133 | 0 |
| CO114700 | 119 | 15 | 133 | 0 |
| BM528126 | 108 | 21 | 128 | 0 |
| CO114700 | 16 | 152 | 167 | 0.63 |
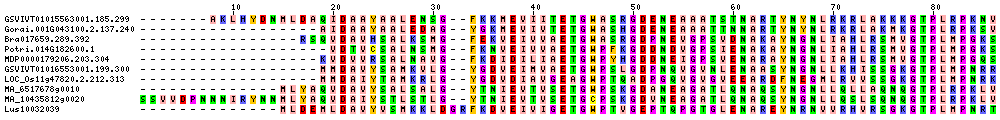
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |