

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_159042g1060 |
| Family | GT9 |
| Protein Properties | Length: 353 Molecular Weight: 38287.4 Isoelectric Point: 9.9576 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT9 | 88 | 329 | 0 |
| QWALAQQFRAAGYGTALVMPRTWKSAIAPTLAGIPQRTGWVGEVRFGLINDWRWGERKMPRFIDKNAALALPANAPLPKQWPVPQLVVPPAEVATWRQAN NLGLGAAVALAPGSVGASKRWTYYAEAARMFAEQGFEVWVVGGPGEKAIATEIVNFAGPRVRDLTGTDLRNGIMAMAAAKTVISNDSGLMHIAAAIGTPA IGIFGPTSPYHWAPLNGLAATIKRATDLPCQPCHKPVCTQND | |||
| Full Sequence |
|---|
| Protein Sequence Length: 353 Download |
| MNINSFLGDI GVTPDPTETS PVLVVPYMWI GDFVRGHTVV RVLKERWPNR PVDLLVTSLC 60 APLVDYMPGV RKGIVWDLPR GQLALKKQWA LAQQFRAAGY GTALVMPRTW KSAIAPTLAG 120 IPQRTGWVGE VRFGLINDWR WGERKMPRFI DKNAALALPA NAPLPKQWPV PQLVVPPAEV 180 ATWRQANNLG LGAAVALAPG SVGASKRWTY YAEAARMFAE QGFEVWVVGG PGEKAIATEI 240 VNFAGPRVRD LTGTDLRNGI MAMAAAKTVI SNDSGLMHIA AAIGTPAIGI FGPTSPYHWA 300 PLNGLAATIK RATDLPCQPC HKPVCTQNDH HCMRDITASE VVETAQRVIA SAR 360 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam01075 | Glyco_transf_9 | 2.0e-18 | 89 | 325 | 245 | + Glycosyltransferase family 9 (heptosyltransferase). Members of this family belong to glycosyltransferase family 9. Lipopolysaccharide is a major component of the outer leaflet of the outer membrane in Gram-negative bacteria. It is composed of three domains; lipid A, Core oligosaccharide and the O-antigen. All of these enzymes transfer heptose to the lipopolysaccharide core. | ||
| COG0859 | RfaF | 2.0e-35 | 22 | 350 | 335 | + ADP-heptose:LPS heptosyltransferase [Cell envelope biogenesis, outer membrane] | ||
| PRK10916 | PRK10916 | 2.0e-37 | 22 | 351 | 347 | + ADP-heptose:LPS heptosyltransferase II; Provisional | ||
| cd03789 | GT1_LPS_heptosyltransferase | 8.0e-62 | 21 | 347 | 331 | + Lipopolysaccharide heptosyltransferase is involved in the biosynthesis of lipooligosaccharide (LOS). Lipopolysaccharide (LPS) is a major component of the outer membrane of gram-negative bacteria. LPS heptosyltransferase transfers heptose molecules from ADP-heptose to 3-deoxy-D-manno-octulosonic acid (KDO), a part of the inner core component of LPS. This family belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| TIGR02195 | heptsyl_trn_II | 2.0e-64 | 22 | 350 | 335 | + lipopolysaccharide heptosyltransferase II. This family consists of examples of ADP-heptose:LPS heptosyltransferase II, an enzyme of LPS inner core region biosynthesis. LPS, composed of lipid A, a core region, and O antigen, is found in the outer membrane of Gram-negative bacteria [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides]. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | YP_001241530.1 | 0 | 1 | 352 | 1 | 348 | ADP-heptose--LPS heptosyltransferase II [Bradyrhizobium sp. BTAi1] |
| RefSeq | YP_002288330.1 | 0 | 1 | 351 | 1 | 350 | lipopolysaccharide heptosyltransferase II [Oligotropha carboxidovorans OM5] |
| RefSeq | YP_317683.1 | 0 | 15 | 352 | 13 | 350 | lipopolysaccharide heptosyltransferase II [Nitrobacter winogradskyi Nb-255] |
| RefSeq | YP_576582.1 | 0 | 12 | 352 | 11 | 351 | lipopolysaccharide heptosyltransferase II [Nitrobacter hamburgensis X14] |
| RefSeq | ZP_01045978.1 | 0 | 12 | 353 | 11 | 352 | lipopolysaccharide heptosyltransferase II [Nitrobacter sp. Nb-311A] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1psw_A | 0 | 22 | 343 | 3 | 337 | A Chain A, Structure Of E. Coli Adp-Heptose Lps Heptosyltransferase Ii |
| PDB | 3tov_B | 0.000000004 | 22 | 353 | 11 | 349 | A Chain A, The Crystal Structure Of The Glycosyl Transferase Family 9 From Veillonella Parvula Dsm 2008 |
| PDB | 3tov_A | 0.000000004 | 22 | 353 | 11 | 349 | A Chain A, The Crystal Structure Of The Glycosyl Transferase Family 9 From Veillonella Parvula Dsm 2008 |
| PDB | 2h1h_B | 0.003 | 205 | 296 | 191 | 284 | A Chain A, The Crystal Structure Of The Glycosyl Transferase Family 9 From Veillonella Parvula Dsm 2008 |
| PDB | 2h1h_A | 0.003 | 205 | 296 | 191 | 284 | A Chain A, The Crystal Structure Of The Glycosyl Transferase Family 9 From Veillonella Parvula Dsm 2008 |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
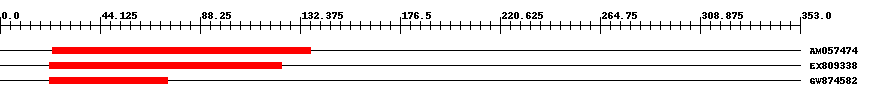 | ||||
| Hit | Length | Start | End | EValue |
| AM057474 | 115 | 23 | 137 | 2e-18 |
| EX809338 | 103 | 22 | 124 | 0.000000000001 |
| GW874582 | 53 | 22 | 74 | 0.061 |
| Orthologous Group | |||||
|---|---|---|---|---|---|
| Species | ID | ||||
| Picea abies | MA_159042g1050 | ||||
| Ricinus communis | 27698.m000066 | ||||
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |