

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000251883 |
| Family | GT1 |
| Protein Properties | Length: 431 Molecular Weight: 47853.8 Isoelectric Point: 5.8658 |
| Chromosome | Chromosome/Scaffold: 015653244 Start: 26884 End: 28345 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT1 | 216 | 396 | 1.4e-37 |
| ESVIYVSFGSGGTLSFEQMTELAWGLELSQQRFIWVIRPPTLTADGAFFTGGSSSDDKNPIKYLPNGFLDRTKDIGLVIPLWAPQVDILSHPAVGGFLSH CGWNSTLESITNGVPMITWPLYAEQRMNATLLTEELGVAVRSKIPPWKKVVEREEIEEMVRKIMEDKEGFALRDRVKKIKE | |||
| Full Sequence |
|---|
| Protein Sequence Length: 431 Download |
| MEMITSSKQH AALLCSPGMG HLIPVLELGK RLVTHXNFTV TIFVISSHTS DAESQLINTA 60 TNPKLCRVVD LPPPDISGLV GPDAAVVTIL AVMMREVRHA FRSALRAMEF CPTVLIVDLF 120 GTESFSIADE FGIPKYIYIP SSAWFLSFMI YGPTLDLEVK GEFTQQKEPI KIPGCRSILP 180 QLDVVDSFRN RTHQQYFDFV EICSQTSKGD GVLLNESVIY VSFGSGGTLS FEQMTELAWG 240 LELSQQRFIW VIRPPTLTAD GAFFTGGSSS DDKNPIKYLP NGFLDRTKDI GLVIPLWAPQ 300 VDILSHPAVG GFLSHCGWNS TLESITNGVP MITWPLYAEQ RMNATLLTEE LGVAVRSKIP 360 PWKKVVEREE IEEMVRKIME DKEGFALRDR VKKIKESGXK ALEKGGSSYN ALSQFAKHSE 420 LMCMKFVNGG E |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02554 | PLN02554 | 3.0e-47 | 217 | 415 | 210 | + UDP-glycosyltransferase family protein | ||
| PLN03015 | PLN03015 | 3.0e-54 | 10 | 215 | 208 | + UDP-glucosyl transferase | ||
| PLN03015 | PLN03015 | 5.0e-69 | 217 | 422 | 208 | + UDP-glucosyl transferase | ||
| PLN00164 | PLN00164 | 2.0e-73 | 13 | 416 | 481 | + glucosyltransferase; Provisional | ||
| PLN02992 | PLN02992 | 2.0e-132 | 7 | 420 | 475 | + coniferyl-alcohol glucosyltransferase | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI25658.1 | 0 | 4 | 416 | 1 | 440 | unnamed protein product [Vitis vinifera] |
| Swiss-Prot | Q40287 | 0 | 6 | 420 | 7 | 474 | UFOG5_MANES RecName: Full=Anthocyanidin 3-O-glucosyltransferase 5; AltName: Full=Flavonol 3-O-glucosyltransferase 5; AltName: Full=UDP-glucose flavonoid 3-O-glucosyltransferase 5 |
| RefSeq | XP_002264323.1 | 0 | 4 | 416 | 1 | 462 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002273356.1 | 0 | 7 | 422 | 4 | 472 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310202.1 | 0 | 1 | 421 | 1 | 476 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2vg8_A | 0 | 3 | 416 | 1 | 464 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2vch_A | 0 | 3 | 416 | 1 | 464 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2vce_A | 0 | 3 | 416 | 1 | 464 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 2acw_B | 8e-40 | 16 | 408 | 17 | 450 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| PDB | 2acw_A | 8e-40 | 16 | 408 | 17 | 450 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt71g1 Complexed With Udp-Glucose |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
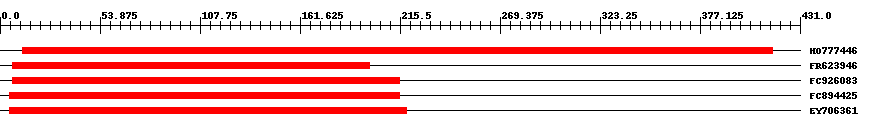 | ||||
| Hit | Length | Start | End | EValue |
| HO777446 | 458 | 12 | 416 | 0 |
| FR623946 | 193 | 7 | 199 | 0 |
| FC926083 | 209 | 7 | 215 | 0 |
| FC894425 | 211 | 5 | 215 | 0 |
| EY706361 | 215 | 5 | 219 | 0 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |