

| Basic Information | |
|---|---|
| Species | Capsella rubella |
| Cazyme ID | Carubv10008071m |
| Family | CBM57 |
| Protein Properties | Length: 1951 Molecular Weight: 214410 Isoelectric Point: 7.3395 |
| Chromosome | Chromosome/Scaffold: 1 Start: 19225321 End: 19239255 |
| Description | Leucine-rich repeat transmembrane protein kinase |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM57 | 409 | 563 | 3.4e-23 |
| INCGGPQIAAVSGEVFEREDEDLGAASFVVSDVQRWAATSVGAFSRSISNIWVVNTLDSDLFRSARQSSSSLRYYGLGLENGVYTVTLQFAEIQIIGSNT WKGLGRRIFDIYVQGRLVEKDFNIRRIVGGDATVGGVQRDYQANVSENYLEIHLL | |||
| CBM57 | 1354 | 1508 | 3.9e-24 |
| SINCGGPQIRSVSGEVFERDDKDLGQASFVVSDIERWAVSSVGLFSGSSNNIWVINALDSELFRSARHSSSSLRYYGLGLENGGYTITLQFAEILIFGSK TWRSLGRRLFDIYVQGRLVEKDFDIRRTAGDTAVRAVRREYKANVSENYIEIHLF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1951 Download |
| MTTCLLLTVW FFLCISGSVY RIRAQNQTVA TTHPDEARAL NSIFAAWRIR APREWNISGE 60 LCSGAATDER ITMEDKNHNP LIKCDCYFEN STICRITALK VFAKDVSGPI PPLLWTLTYL 120 TDLNLAQNYL TGSLSPAIGN LTQLEWLAIG INALSGPFPK EIGLLTNLKS LGIGLNNFTG 180 PIPAEIGNCT RLLKIYLGNS GLGGEIPSSF ANLIEMQEAC INDVDVTGRI PEFIGTWTKL 240 LILKLHGTGL SGPLPSSFSN LTSLTDLSVG DISNGGSSLE FIKGMKSLNI LTFRNSNLTG 300 TIPSNIGEYS KLKHVDLSFN KLHGPIPASL FNISGLTYLF LGNNTFNGSL PTQKSQTLSN 360 IDVSYNDLSG SLPSWVSLPN LKLVLPGLKC LQKNFPCNRG KGIYSDFWIN CGGPQIAAVS 420 GEVFEREDED LGAASFVVSD VQRWAATSVG AFSRSISNIW VVNTLDSDLF RSARQSSSSL 480 RYYGLGLENG VYTVTLQFAE IQIIGSNTWK GLGRRIFDIY VQGRLVEKDF NIRRIVGGDA 540 TVGGVQRDYQ ANVSENYLEI HLLWAGKGSI GIPVMGTYGP LVSAVSAKPD FIPTVANRPP 600 SKKKYKTGTV VGAGLGLGLL SIFFAGVVTF IIRKSKKRYT VDKELLSMEI KPYTFTYSEL 660 KSATQDFNTS NKLGEGGFGS VYKGTLSDGR EIAVKVLSVG SQHGKGQFVA EIVTISTVLH 720 RNLVKLYGCC YEGDHRLLVY EYLPNGSLDH ALFGGEKTLH LDWSTRFEIC LGVARGLAYL 780 HEEASDRIVH RDVKASNILL DSKLLPKVSD FGLAKLYDDS NTHMSTKVAG TIGYLSPEYA 840 MRGQLSEKTD VYAFGVVVLE LVSGRKNYDT NLDDEKKYLL DWAWNQHEQS REGDLIDHRL 900 TEFNMEEAKR MIGIALLCIQ ASYGLRPPMS RVVAMLSGDV EVSDVTSKPG YLTDWTFDET 960 SSSSFGTFQT KDTTASSSSS TMFVTPREAD TARALNSIFA AWRIRAPREW NISGELCSGA 1020 AIDERITIDD KNHNPFIKCD CNFENSTICH ITALKVFQKD VVGAIPPQLW TLTYLINLNL 1080 AQNYLTGSIP PAIGYLTRME WLTFGINALS GPFPKEIGLL TELKSLGIGV NNFSGSIPAE 1140 IGNCTKLMKI YLGVSGLSGE IPLSFANLVE LQEAYITDMD ITGRIPKFIG TWTKLTILKI 1200 VGTGLSGPIP SSFSNLTSLK ELVFRNSNLT GTIASNIGEY SSLQQVDLSF NKLHGPIPAS 1260 LFNLNQLTHL FLGNNTFNGS LPTQKSQALT NIDVSYNNLS GNLPSWVSLP NLKLNLVVNN 1320 FTLEGLDKRV LPGLKCLQKN FPCNRGKGIY SDFSINCGGP QIRSVSGEVF ERDDKDLGQA 1380 SFVVSDIERW AVSSVGLFSG SSNNIWVINA LDSELFRSAR HSSSSLRYYG LGLENGGYTI 1440 TLQFAEILIF GSKTWRSLGR RLFDIYVQGR LVEKDFDIRR TAGDTAVRAV RREYKANVSE 1500 NYIEIHLFWA GKGSFSVPVI GTFGPLISAV TAKPDFTPTV TNRPPSKKKN RTGIIVGVTT 1560 SIGLLSIFFA GVIIFIIRKS KKRYTNDEEL LFMDVKPYTF AYSELKSATQ DFDPSNKLGE 1620 GGFGPVYKGT LSDGREIAVK ELSVGSRQGK GQFVAEIVTI STVLHRNLVK LYGCCYEGDH 1680 RLLVYEYLPN GSLDHALFGH GGEKTLHLDW STRFEICMGV ARGLAYLHEE ASVCIVHRDV 1740 KASNILLDSK LLPKVSDFGL AKLYDDKKTH ISTRVAGTIG YLAPEYAMRG YLTEKTDVYA 1800 FGVVVLELVS GRPNSDVSLN DEKKFLLEWA WNLHEKNSEI ELIDPDLTEI NMDEVQRMIG 1860 IALLCIQSSY GLRPPMSRVV AMLSGDIEVT DATSKPGYLT DWRINDTSSS SFSAFQATDT 1920 STSWASSTRF VTPKDSDFKP MLGLKINEGR * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
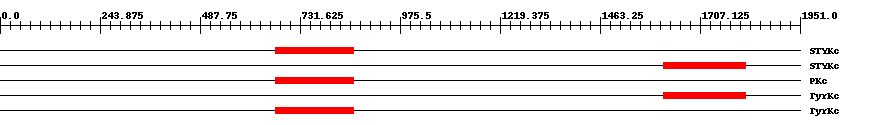 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| smart00221 | STYKc | 2.0e-46 | 672 | 863 | 198 | + Protein kinase; unclassified specificity. Phosphotransferases. The specificity of this class of kinases can not be predicted. Possible dual-specificity Ser/Thr/Tyr kinase. | ||
| smart00221 | STYKc | 6.0e-47 | 1617 | 1817 | 214 | + Protein kinase; unclassified specificity. Phosphotransferases. The specificity of this class of kinases can not be predicted. Possible dual-specificity Ser/Thr/Tyr kinase. | ||
| cd00180 | PKc | 6.0e-47 | 673 | 861 | 193 | + Catalytic domain of Protein Kinases. Protein Kinases (PKs), catalytic (c) domain. PKs catalyze the transfer of the gamma-phosphoryl group from ATP to serine/threonine or tyrosine residues on protein substrates. The PK family is part of a larger superfamily that includes the catalytic domains of RIO kinases, aminoglycoside phosphotransferase, choline kinase, phosphoinositide 3-kinase (PI3K), and actin-fragmin kinase. PKs make up a large family of serine/threonine kinases, protein tyrosine kinases (PTKs), and dual-specificity PKs that phosphorylate both serine/threonine and tyrosine residues of target proteins. Majority of protein phosphorylation, about 95%, occurs on serine residues while only 1% occurs on tyrosine residues. Protein phosphorylation is a mechanism by which a wide variety of cellular proteins, such as enzymes and membrane channels, are reversibly regulated in response to certain stimuli. PKs often function as components of signal transduction pathways in which one kinase activates a second kinase, which in turn, may act on other kinases; this sequential action transmits a signal from the cell surface to target proteins, which results in cellular responses. The PK family is one of the largest known protein families with more than 100 homologous yeast enzymes and 550 human proteins. A fraction of PK family members are pseudokinases that lack crucial residues for catalytic activity. The mutiplicity of kinases allows for specific regulation according to substrate, tissue distribution, and cellular localization. PKs regulate many cellular processes including proliferation, division, differentiation, motility, survival, metabolism, cell-cycle progression, cytoskeletal rearrangement, immunity, and neuronal functions. Many kinases are implicated in the development of various human diseases including different types of cancer. | ||
| smart00219 | TyrKc | 5.0e-48 | 1617 | 1817 | 214 | + Tyrosine kinase, catalytic domain. Phosphotransferases. Tyrosine-specific kinase subfamily. | ||
| smart00219 | TyrKc | 4.0e-48 | 672 | 863 | 198 | + Tyrosine kinase, catalytic domain. Phosphotransferases. Tyrosine-specific kinase subfamily. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004672 | protein kinase activity |
| GO:0005524 | ATP binding |
| GO:0006468 | protein phosphorylation |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAF02838.1 | 0 | 991 | 1950 | 14 | 1029 | AC009894_9 Similar to serine/threonine kinases [Arabidopsis thaliana] |
| EMBL | CAJ86064.1 | 0 | 23 | 955 | 846 | 1731 | H0818E11.1 [Oryza sativa (indica cultivar-group)] |
| EMBL | CAJ86064.1 | 0 | 29 | 1928 | 21 | 1751 | H0818E11.1 [Oryza sativa (indica cultivar-group)] |
| RefSeq | NP_176009.1 | 0 | 4 | 997 | 9 | 1026 | leucine-rich repeat family protein / protein kinase family protein [Arabidopsis thaliana] |
| RefSeq | NP_176009.1 | 0 | 990 | 1950 | 40 | 1032 | leucine-rich repeat family protein / protein kinase family protein [Arabidopsis thaliana] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3tl8_H | 0 | 655 | 939 | 28 | 317 | B Chain B, The Avrptob-Bak1 Complex Reveals Two Structurally Similar Kinaseinteracting Domains In A Single Type Iii Effector |
| PDB | 3tl8_H | 0 | 1600 | 1886 | 28 | 317 | B Chain B, The Avrptob-Bak1 Complex Reveals Two Structurally Similar Kinaseinteracting Domains In A Single Type Iii Effector |
| PDB | 3tl8_G | 0 | 655 | 939 | 28 | 317 | B Chain B, The Avrptob-Bak1 Complex Reveals Two Structurally Similar Kinaseinteracting Domains In A Single Type Iii Effector |
| PDB | 3tl8_G | 0 | 1600 | 1886 | 28 | 317 | B Chain B, The Avrptob-Bak1 Complex Reveals Two Structurally Similar Kinaseinteracting Domains In A Single Type Iii Effector |
| PDB | 3tl8_D | 0 | 655 | 939 | 28 | 317 | B Chain B, The Avrptob-Bak1 Complex Reveals Two Structurally Similar Kinaseinteracting Domains In A Single Type Iii Effector |