

| Basic Information | |
|---|---|
| Species | Glycine max |
| Cazyme ID | Glyma13g42560.1 |
| Family | GH35 |
| Protein Properties | Length: 709 Molecular Weight: 80180.6 Isoelectric Point: 7.8979 |
| Chromosome | Chromosome/Scaffold: 13 Start: 42511025 End: 42521598 |
| Description | beta-galactosidase 17 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH35 | 75 | 399 | 0 |
| FWKDGEPFQIIGGDVHYFRVHPEYWEDRLLKAKALGLNTIQTYVPWNLHEPAPGKLVFEGFANIEAFLNLCHKHGLLVMIRPGPYICGEWDWGGFPGWFY SMIPTPKPRSSDPTYLQLVERWWGNLLPKFVPLLYENGGPIIMVQIENEYGSYGDDKEYLHHLITLARGHLGHDVILYTTDGGTRETLEKGTIRGDTIFS AVDFGTGEDPWPIFKLQKEFNAPGKSPPLSAEFYTGWLTHWGEKNAQTDADFTAAALEKILQKNGSAVLYMAHGGTNFGFYNGANTGVDEADYKPDLTSY DYDAPIRESGDVDNSKFNAIRRVIA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 709 Download |
| MRIGLKKKKT TSGAASKLLA MAKKLSSKPT ILFIFVSFML FCAFLPVFAP LPSFSSHHSH 60 RNTVNRKFEI ANDRFWKDGE PFQIIGGDVH YFRVHPEYWE DRLLKAKALG LNTIQTYVPW 120 NLHEPAPGKL VFEGFANIEA FLNLCHKHGL LVMIRPGPYI CGEWDWGGFP GWFYSMIPTP 180 KPRSSDPTYL QLVERWWGNL LPKFVPLLYE NGGPIIMVQI ENEYGSYGDD KEYLHHLITL 240 ARGHLGHDVI LYTTDGGTRE TLEKGTIRGD TIFSAVDFGT GEDPWPIFKL QKEFNAPGKS 300 PPLSAEFYTG WLTHWGEKNA QTDADFTAAA LEKILQKNGS AVLYMAHGGT NFGFYNGANT 360 GVDEADYKPD LTSYDYDAPI RESGDVDNSK FNAIRRVIAR YSSVPLPSIP SNNEKARYGP 420 IHLQREAFVF DMFDFTNSTN VFKSETPMSM EYVGQLFGFV LYVTEYKAKR GGRILFIPKL 480 HDRAQVFISC PSEESGARPT YIGTIERWLN NKVTLPDIKC HSKINLFILV ENMGRVNYGS 540 FIFDRKGILS SVYLDKEQVK GWKMFPIPLH NLNEMSTYNP ITQVAYSAFS GISSFRKKLI 600 YKNGNTSKEP AFYSGHFLID KSSQVKDTFI SFNNWGKGIV FVNDFNIGRY WPLRGPQCNL 660 YVPAPLLKQG DNFLVILELE SPDPELVVHT VDEPDFTCGS SGTSLHQL* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
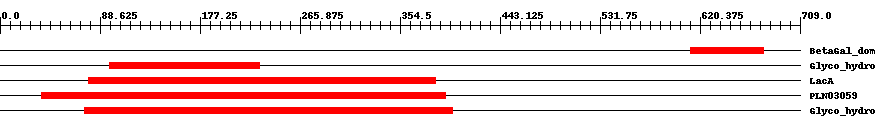 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam13364 | BetaGal_dom4_5 | 0.0004 | 612 | 677 | 70 | + Beta-galactosidase jelly roll domain. This domain is found in beta galactosidase enzymes. It has a jelly roll fold. | ||
| pfam02449 | Glyco_hydro_42 | 8.0e-7 | 97 | 230 | 159 | + Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. | ||
| COG1874 | LacA | 2.0e-32 | 78 | 386 | 343 | + Beta-galactosidase [Carbohydrate transport and metabolism] | ||
| PLN03059 | PLN03059 | 8.0e-40 | 37 | 395 | 379 | + beta-galactosidase; Provisional | ||
| pfam01301 | Glyco_hydro_35 | 2.0e-150 | 75 | 401 | 327 | + Glycosyl hydrolases family 35. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0004565 | beta-galactosidase activity |
| GO:0005975 | carbohydrate metabolic process |
| GO:0009341 | beta-galactosidase complex |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABD28499.1 | 0 | 217 | 708 | 1 | 492 | Galactose-binding like [Medicago truncatula] |
| EMBL | CBI35958.1 | 0 | 9 | 705 | 13 | 707 | unnamed protein product [Vitis vinifera] |
| RefSeq | NP_565051.1 | 0 | 64 | 708 | 59 | 697 | BGAL17 (beta-galactosidase 17); beta-galactosidase/ catalytic/ cation binding [Arabidopsis thaliana] |
| RefSeq | XP_002280228.1 | 0 | 21 | 705 | 1 | 683 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002327549.1 | 0 | 66 | 708 | 1 | 643 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3thd_D | 0 | 61 | 706 | 2 | 629 | A Chain A, Structure And Mechanisim Of Core 2 Beta1,6-N- Acetylglucosaminyltransferase: A Metal-Ion Independent Gt-A Glycosyltransferase |
| PDB | 3thd_C | 0 | 61 | 706 | 2 | 629 | A Chain A, Structure And Mechanisim Of Core 2 Beta1,6-N- Acetylglucosaminyltransferase: A Metal-Ion Independent Gt-A Glycosyltransferase |
| PDB | 3thd_B | 0 | 61 | 706 | 2 | 629 | A Chain A, Structure And Mechanisim Of Core 2 Beta1,6-N- Acetylglucosaminyltransferase: A Metal-Ion Independent Gt-A Glycosyltransferase |
| PDB | 3thd_A | 0 | 61 | 706 | 2 | 629 | A Chain A, Structure And Mechanisim Of Core 2 Beta1,6-N- Acetylglucosaminyltransferase: A Metal-Ion Independent Gt-A Glycosyltransferase |
| PDB | 3thc_D | 0 | 61 | 706 | 2 | 629 | A Chain A, Crystal Structure Of Human Beta-Galactosidase In Complex With Galactose |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| lactose degradation III | BETAGALACTOSID-RXN | EC-3.2.1.23 | β-galactosidase |