

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_10435226g0010 |
| Family | AA1 |
| Protein Properties | Length: 326 Molecular Weight: 36342.2 Isoelectric Point: 7.9745 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA1 | 3 | 326 | 0 |
| PKRISRLCKQHTIITVNGQLPGPTIVVRDGDTVNIKTYNRAQYNATLHWHGVEQLRTSWADGPAYITQCPIPPGGRYTYRFKISGQGGTVWWHAHISWLR ATVHGVFVIYPKRGTLYPFPKYRADIPIVIGEWWNDNPIAVIDEAVLTGGPPNLSDAYTLNGQPGDLFPCSSSETFRLPVERGETYLLRIVNAALNTGHF FKIADHEFTVVAVDACYTKPYKTDVIVISAGQTTDVLLTANQSLGKYYMAARPYNNQAAGNFDNSTTTAILEYIGAKNSTLPILPSFPSYNDTETVTKFN RALRSLASQEYPVDVPRMIDESLI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 326 Download |
| IEPKRISRLC KQHTIITVNG QLPGPTIVVR DGDTVNIKTY NRAQYNATLH WHGVEQLRTS 60 WADGPAYITQ CPIPPGGRYT YRFKISGQGG TVWWHAHISW LRATVHGVFV IYPKRGTLYP 120 FPKYRADIPI VIGEWWNDNP IAVIDEAVLT GGPPNLSDAY TLNGQPGDLF PCSSSETFRL 180 PVERGETYLL RIVNAALNTG HFFKIADHEF TVVAVDACYT KPYKTDVIVI SAGQTTDVLL 240 TANQSLGKYY MAARPYNNQA AGNFDNSTTT AILEYIGAKN STLPILPSFP SYNDTETVTK 300 FNRALRSLAS QEYPVDVPRM IDESLI |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02191 | PLN02191 | 2.0e-34 | 1 | 314 | 344 | + L-ascorbate oxidase | ||
| TIGR03388 | ascorbase | 1.0e-43 | 10 | 307 | 326 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| pfam00394 | Cu-oxidase | 9.0e-44 | 127 | 278 | 156 | + Multicopper oxidase. Many of the proteins in this family contain multiple similar copies of this plastocyanin-like domain. | ||
| pfam07732 | Cu-oxidase_3 | 3.0e-50 | 1 | 116 | 118 | + Multicopper oxidase. This entry contains many divergent copper oxidase-like domains that are not recognised by the pfam00394 model. | ||
| TIGR03389 | laccase | 1.0e-172 | 1 | 326 | 326 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK37824.1 | 0 | 6 | 326 | 47 | 369 | AF132120_1 laccase [Pinus taeda] |
| GenBank | AAK37826.1 | 0 | 1 | 326 | 37 | 363 | AF132122_1 laccase [Pinus taeda] |
| GenBank | ABR17539.1 | 0 | 1 | 326 | 36 | 366 | unknown [Picea sitchensis] |
| GenBank | ACN40347.1 | 0 | 1 | 326 | 38 | 363 | unknown [Picea sitchensis] |
| RefSeq | XP_002277722.1 | 0 | 1 | 326 | 41 | 367 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pxl_A | 1e-37 | 1 | 314 | 15 | 314 | A Chain A, Type-2 Cu-Depleted Fungus Laccase From Trametes Hirsuta |
| PDB | 3v9c_A | 1e-37 | 1 | 314 | 15 | 314 | A Chain A, Type-2 Cu-Depleted Fungus Laccase From Trametes Hirsuta |
| PDB | 3fpx_A | 1e-37 | 1 | 314 | 15 | 314 | A Chain A, Native Fungus Laccase From Trametes Hirsuta |
| PDB | 3div_A | 4e-35 | 16 | 314 | 25 | 314 | A Chain A, Crystal Structure Of Laccase From Cerrena Maxima At 1.76a Resolution |
| PDB | 2h5u_A | 5e-35 | 16 | 278 | 25 | 284 | A Chain A, Crystal Structure Of Laccase From Cerrena Maxima At 1.9a Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| CO489402 | 235 | 92 | 326 | 0 |
| GT741923 | 275 | 1 | 275 | 0 |
| DV996223 | 275 | 1 | 275 | 0 |
| CO484010 | 271 | 56 | 326 | 0 |
| GE481798 | 271 | 56 | 326 | 0 |
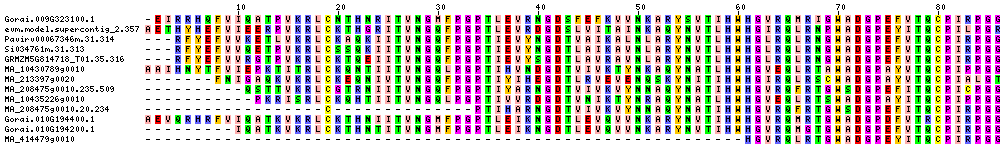
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |