

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_414479g0010 |
| Family | AA1 |
| Protein Properties | Length: 458 Molecular Weight: 50211.6 Isoelectric Point: 6.3829 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA1 | 1 | 188 | 0 |
| HGVRQLRTGWADGPEYITQCPIRPGGSYTYRFNITCQEGTLWWHAHSSWLRATVYGALIIYPSLGTSYPFARPDRQIPIVLGEWWNRNPIDVVNQATTTG AAPSVSDAFTINGQPGGLYPCSSSDTFRASVKAGETNLLRIINAGMNTDLFFSVANHTMTVVAVDASYTKPLQTNVLMLGPGQTTDTV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 458 Download |
| HGVRQLRTGW ADGPEYITQC PIRPGGSYTY RFNITCQEGT LWWHAHSSWL RATVYGALII 60 YPSLGTSYPF ARPDRQIPIV LGEWWNRNPI DVVNQATTTG AAPSVSDAFT INGQPGGLYP 120 CSSSDTFRAS VKAGETNLLR IINAGMNTDL FFSVANHTMT VVAVDASYTK PLQTNVLMLG 180 PGQTTDTVTS FADGLRSLAT QDHPVFVPQS VDENLFYTVG LGLISCPGQS CGSPNGSRMA 240 ASMNNISFVL PTSFSILQAQ QLGKKGVFTT DFPDNPPVQF DYTAQNISRG LWSPVKDTRV 300 KVLTYNTTVQ VILQGTNIFA GESHPIHLHG YDFYIVGAGF GNYNSQTDPQ KFNLVDESHP 360 IHLHGYDFYI VGAGFGNYNS QTDPQKFNLV DPPMRNTVNV PVNGWAAIRF VADNPGAWLM 420 HCHLDVHATW GLAMVFVVNN GPDYLLSLEP PPRDLPLC |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR03388 | ascorbase | 3.0e-31 | 242 | 436 | 200 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| pfam07731 | Cu-oxidase_2 | 3.0e-38 | 271 | 442 | 176 | + Multicopper oxidase. This entry contains many divergent copper oxidase-like domains that are not recognised by the pfam00394 model. | ||
| PLN02191 | PLN02191 | 6.0e-40 | 1 | 437 | 507 | + L-ascorbate oxidase | ||
| PLN02604 | PLN02604 | 3.0e-49 | 1 | 436 | 509 | + oxidoreductase | ||
| TIGR03389 | laccase | 0 | 1 | 458 | 516 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK37824.1 | 0 | 1 | 458 | 93 | 576 | AF132120_1 laccase [Pinus taeda] |
| GenBank | AAK37826.1 | 0 | 1 | 458 | 88 | 570 | AF132122_1 laccase [Pinus taeda] |
| GenBank | ABR17528.1 | 0 | 1 | 458 | 89 | 570 | unknown [Picea sitchensis] |
| GenBank | ABR17539.1 | 0 | 1 | 458 | 87 | 573 | unknown [Picea sitchensis] |
| RefSeq | NP_196158.1 | 0 | 1 | 458 | 84 | 565 | LAC12 (laccase 12); laccase [Arabidopsis thaliana] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1asq_B | 2e-37 | 1 | 437 | 62 | 522 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 1asq_A | 2e-37 | 1 | 437 | 62 | 522 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 1asp_B | 2e-37 | 1 | 437 | 62 | 522 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 1asp_A | 2e-37 | 1 | 437 | 62 | 522 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 1aso_B | 2e-37 | 1 | 437 | 62 | 522 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR102424 | 274 | 185 | 458 | 0 |
| EX351374 | 273 | 186 | 458 | 0 |
| DR023255 | 274 | 185 | 458 | 0 |
| DV977806 | 274 | 185 | 458 | 0 |
| DV971407 | 270 | 189 | 458 | 0 |
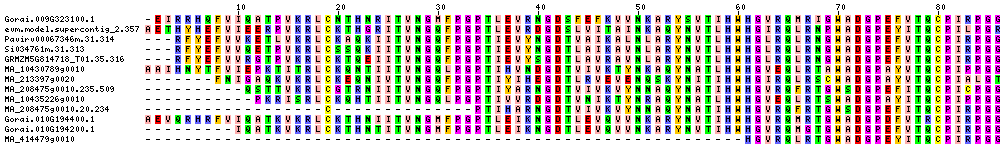
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |